|
1
|
Felch KL, Crider JD, Bhattacharjee D, Huhn C, Wilson M, Bengtén E. TLR7 in channel catfish (Ictalurus punctatus) is expressed in the endolysosome and is stimulated by synthetic ssRNA analogs, imiquimod, and resiquimod. DEVELOPMENTAL AND COMPARATIVE IMMUNOLOGY 2024; 157:105197. [PMID: 38763479 PMCID: PMC11234115 DOI: 10.1016/j.dci.2024.105197] [Citation(s) in RCA: 0] [Impact Index Per Article: 0] [Reference Citation Analysis] [Abstract] [MESH Headings] [Grants] [Track Full Text] [Subscribe] [Scholar Register] [Received: 02/27/2024] [Revised: 05/16/2024] [Accepted: 05/16/2024] [Indexed: 05/21/2024]
Abstract
Toll-like receptors (TLRs) are pivotal pattern recognition receptors (PRRs) and key mediators of innate immunity. Despite the significance of channel catfish (Ictalurus punctatus) in comparative immunology and aquaculture, its 20 TLR genes remain largely functionally uncharacterized. In this study, our aim was to determine the catfish TLR7 agonists, signaling potential, and cellular localization. Using a mammalian reporter system, we identified imiquimod and resiquimod, typical ssRNA analogs, as potent catfish TLR7 agonists. Notably, unlike grass carp TLR7, catfish TLR7 lacks the ability to respond to poly (I:C). Confocal microscopy revealed predominant catfish TLR7 expression in lysosomes, co-localizing with the endosomal chaperone protein, UNC93B1. Furthermore, imiquimod stimulation elicited robust IFNb transcription in peripheral blood leukocytes isolated from adult catfish. These findings underscore the conservation of TLR7 signaling in catfish, reminiscent of mammalian TLR7 responses. Our study sheds light on the functional aspects of catfish TLR7 and contributes to a better understanding of its role in immune defense mechanisms.
Collapse
Affiliation(s)
- Kristianna L Felch
- Department of Cell and Molecular Biology, University of Mississippi Medical Center, 2500 North State Street, 39216, Jackson, MS, USA.
| | - Jonathan D Crider
- Department of Cell and Molecular Biology, University of Mississippi Medical Center, 2500 North State Street, 39216, Jackson, MS, USA; Department of Biology, Belmont University, 1900 Belmont Blvd, 37212, Nashville, TN, USA.
| | - Debduti Bhattacharjee
- Department of Cell and Molecular Biology, University of Mississippi Medical Center, 2500 North State Street, 39216, Jackson, MS, USA.
| | - Cameron Huhn
- Department of Cell and Molecular Biology, University of Mississippi Medical Center, 2500 North State Street, 39216, Jackson, MS, USA.
| | - Melanie Wilson
- Department of Cell and Molecular Biology, University of Mississippi Medical Center, 2500 North State Street, 39216, Jackson, MS, USA; Center for Immunology and Microbial Research, University of Mississippi Medical Center, 2500 North State Street, 39216, Jackson, MS, USA.
| | - Eva Bengtén
- Department of Cell and Molecular Biology, University of Mississippi Medical Center, 2500 North State Street, 39216, Jackson, MS, USA; Center for Immunology and Microbial Research, University of Mississippi Medical Center, 2500 North State Street, 39216, Jackson, MS, USA.
| |
Collapse
|
|
2
|
Harman RM, Sipka A, Oxford KA, Oliveira L, Huntimer L, Nydam DV, Van de Walle GR. The mammosphere-derived epithelial cell secretome modulates neutrophil functions in the bovine model. Front Immunol 2024; 15:1367432. [PMID: 38994364 PMCID: PMC11236729 DOI: 10.3389/fimmu.2024.1367432] [Citation(s) in RCA: 0] [Impact Index Per Article: 0] [Reference Citation Analysis] [Abstract] [Key Words] [MESH Headings] [Track Full Text] [Figures] [Journal Information] [Subscribe] [Scholar Register] [Received: 01/08/2024] [Accepted: 06/17/2024] [Indexed: 07/13/2024] Open
Abstract
Background Innovative therapies against bacterial infections are needed. One approach is to focus on host-directed immunotherapy (HDT), with treatments that exploit natural processes of the host immune system. The goals of this type of therapy are to stimulate protective immunity while minimizing inflammation-induced tissue damage. We use non-traditional large animal models to explore the potential of the mammosphere-derived epithelial cell (MDEC) secretome, consisting of all bioactive factors released by the cells, to modulate host immune functions. MDEC cultures are enriched for mammary stem and progenitor cells and can be generated from virtually any mammal. We previously demonstrated that the bovine MDEC secretome, collected and delivered as conditioned medium (CM), inhibits the growth of bacteria in vitro and stimulates functions related to tissue repair in cultured endothelial and epithelial cells. Methods The immunomodulatory effects of the bovine MDEC secretome on bovine neutrophils, an innate immune cell type critical for resolving bacterial infections, were determined in vitro using functional assays. The effects of MDEC CM on neutrophil molecular pathways were explored by evaluating the production of specific cytokines by neutrophils and examining global gene expression patterns in MDEC CM-treated neutrophils. Enzyme linked immunosorbent assays were used to determine the concentrations of select proteins in MDEC CM and siRNAs were used to reduce the expression of specific MDEC-secreted proteins, allowing for the identification of bioactive factors modulating neutrophil functions. Results Neutrophils exposed to MDEC secretome exhibited increased chemotaxis and phagocytosis and decreased intracellular reactive oxygen species and extracellular trap formation, when compared to neutrophils exposed to control medium. C-X-C motif chemokine 6, superoxide dismutase, peroxiredoxin-2, and catalase, each present in the bovine MDEC secretome, were found to modulate neutrophil functions. Conclusion The MDEC secretome administered to treat bacterial infections may increase neutrophil recruitment to the site of infection, stimulate pathogen phagocytosis by neutrophils, and reduce neutrophil-produced ROS accumulation. As a result, pathogen clearance might be improved and local inflammation and tissue damage reduced.
Collapse
Affiliation(s)
- Rebecca M. Harman
- Baker Institute for Animal Health, College of Veterinary Medicine, Cornell University, Ithaca, NY, United States
| | - Anja Sipka
- Department of Population Medicine and Diagnostic Sciences, College of Veterinary Medicine, Cornell University, Ithaca, NY, United States
| | - Kelly A. Oxford
- Baker Institute for Animal Health, College of Veterinary Medicine, Cornell University, Ithaca, NY, United States
| | | | | | - Daryl V. Nydam
- Department of Public and Ecosystem Health, Cornell University, Ithaca, NY, United States
| | - Gerlinde R. Van de Walle
- Baker Institute for Animal Health, College of Veterinary Medicine, Cornell University, Ithaca, NY, United States
| |
Collapse
|
|
3
|
Pu Z, Wang W, Xie H, Wang W. Apolipoprotein C3 (ApoC3) facilitates NLRP3 mediated pyroptosis of macrophages through mitochondrial damage by accelerating of the interaction between SCIMP and SYK pathway in acute lung injury. Int Immunopharmacol 2024; 128:111537. [PMID: 38232538 DOI: 10.1016/j.intimp.2024.111537] [Citation(s) in RCA: 0] [Impact Index Per Article: 0] [Reference Citation Analysis] [Abstract] [Key Words] [MESH Headings] [Track Full Text] [Journal Information] [Subscribe] [Scholar Register] [Received: 09/03/2023] [Revised: 12/25/2023] [Accepted: 01/10/2024] [Indexed: 01/19/2024]
Abstract
Respiratory failure caused by severe acute lung injury (ALI) is the main cause of mortality in patients with COVID-19.This study aimed to investigate the effects and underlying biological mechanism of Apolipoprotein C3 (ApoC3) in ALI. To establish an in vivo model, C57BL/6 mice were exposed by lipopolysaccharide (LPS). For the in vitro model, murine bone marrow-derived macrophages (BMDMs) or RAW264.7 cells were stimulated with LPS + adenosine triphosphate (ATP). Serum levels of ApoC3 were found to be upregulated in patients with COVID-19 or pneumonia-induced ALI. Inhibition of ApoC3 reduced lung injury in an ALI model, while overexpression of ApoC3 promoted lung injury. ApoC3 induced mitochondrial damage-mediated pyroptosis in ALI through the activation of the NOD-like receptorprotein 3 (NLRP3) inflammasome. ApoC3 recombinant protein significantly increased SCIMP expression in the lung tissue of mice models with ALI. ApoC3 also facilitated the interaction between the SLP adapter and CSK-interacting membrane protein (SCIMP) protein and Spleen tyrosine kinase (SYK) protein in the ALI model. Moreover, ApoC3 accelerated calcium-dependent reactive oxygen species (ROS) production in the ALI model. The effects of ApoC3 on pyroptosis were mitigated by the use of a pyroptosis inhibitor or an ROS inhibitor in the ALI model. Furthermore, ApoC3 activated the expression of SYK, which in turn induced NLRP3 inflammasome-regulated pyroptosis in the ALI model. METTL3 was found to mediate the m6A mRNA expression of ApoC3. Overall, our study highlights the crucial role of ApoC3 in promoting macrophage pyroptosis in ALI through calcium-dependent ROS production and NLRP3 inflammasome activation via the SCIMP-SYK pathway, providing a potential therapeutic strategy for ALI and other inflammatory diseases.
Collapse
Affiliation(s)
- Zhichen Pu
- Drug Clinical Evaluation, Yijishan Hospital of Wannan Medical College, Wuhu, Anhui 241001, China; Key Laboratory of Non-coding RNA Transformation Research of Anhui Higher Education Institution, Wannan Medical College, Wuhu 241001, China; State Key Laboratory of Natural Medicines, Key Lab of Drug Metabolism and Pharmacokinetics, China Pharmaceutical University, Tongjiaxiang 24, Nanjing 210009, China
| | - Wenhui Wang
- Drug Clinical Evaluation, Yijishan Hospital of Wannan Medical College, Wuhu, Anhui 241001, China
| | - Haitang Xie
- Drug Clinical Evaluation, Yijishan Hospital of Wannan Medical College, Wuhu, Anhui 241001, China.
| | - Wusan Wang
- Department of Pharmacology, Wannan Medical College, Wuhu, Anhui 241001, China.
| |
Collapse
|
|
4
|
Pei X, Liu L, Wang J, Guo C, Li Q, Li J, Ren Q, Ma R, Zheng Y, Zhang Y, Liu L, Zheng D, Wang P, Jiang P, Feng X, Jiang E, Wang Y, Feng S. Exosomal secreted SCIMP regulates communication between macrophages and neutrophils in pneumonia. Nat Commun 2024; 15:691. [PMID: 38263143 PMCID: PMC10805922 DOI: 10.1038/s41467-024-44714-4] [Citation(s) in RCA: 0] [Impact Index Per Article: 0] [Reference Citation Analysis] [Abstract] [Key Words] [MESH Headings] [Grants] [Track Full Text] [Journal Information] [Subscribe] [Scholar Register] [Received: 04/11/2023] [Accepted: 01/02/2024] [Indexed: 01/25/2024] Open
Abstract
In pneumonia, the deficient or delayed pathogen clearance can lead to pathogen proliferation and subsequent overactive immune responses, inducing acute lung injury (ALI). While screening human genome coding genes using our peripheral blood cell chemotactic platform, we unexpectedly find SLP adaptor and CSK interacting membrane protein (SCIMP), a protein with neutrophil chemotactic activity secreted during ALI. However, the specific role of SCIMP in ALI remains unclear. In this study, we investigate the secretion of SCIMP in exosomes (SCIMPexo) by macrophages after bacterial stimulation, both in vitro and in vivo. We observe a significant increase in the levels of SCIMPexo in bronchoalveolar lavage fluid and serum of pneumonia patients. We also find that bronchial perfusion with SCIMPexo or SCIMP N-terminal peptides increases the survival rate of the ALI model. This occurs due to the chemoattraction and activation of peripheral neutrophils dependent on formyl peptide receptor 1/2 (FPR1/2). Conversely, exosome suppressors and FPR1/2 antagonists decrease the survival rate in the lethal ALI model. Scimp-deficient and Fpr1/2-deficient mice also have lower survival rates and shorter survival times than wild-type mice. However, bronchial perfusion of SCIMP rescues Scimp-deficient mice but not Fpr1/2-deficient mice. Collectively, our findings suggest that the macrophage-SCIMP-FPRs-neutrophil axis plays a vital role in the innate immune process underlying ALI.
Collapse
Affiliation(s)
- Xiaolei Pei
- State Key Laboratory of Experimental Hematology, National Clinical Research Center for Blood Diseases, Haihe Laboratory of Cell Ecosystem, Hematopoietic Stem Cell Transplantation Center, Institute of Hematology & Blood Diseases Hospital, Chinese Academy of Medical Sciences & Peking Union Medical College, Tianjin, 300020, P. R. China.
- Tianjin Institutes of Health Science, Tianjin, 301600, P. R. China.
| | - Li Liu
- State Key Laboratory of Experimental Hematology, National Clinical Research Center for Blood Diseases, Haihe Laboratory of Cell Ecosystem, Hematopoietic Stem Cell Transplantation Center, Institute of Hematology & Blood Diseases Hospital, Chinese Academy of Medical Sciences & Peking Union Medical College, Tianjin, 300020, P. R. China
- Tianjin Institutes of Health Science, Tianjin, 301600, P. R. China
| | - Jieru Wang
- State Key Laboratory of Experimental Hematology, National Clinical Research Center for Blood Diseases, Haihe Laboratory of Cell Ecosystem, Hematopoietic Stem Cell Transplantation Center, Institute of Hematology & Blood Diseases Hospital, Chinese Academy of Medical Sciences & Peking Union Medical College, Tianjin, 300020, P. R. China
- Tianjin Institutes of Health Science, Tianjin, 301600, P. R. China
| | - Changyuan Guo
- Department of Immunology, School of Basic Medical Sciences and NHC Key Laboratory of Medical Immunology, Peking University, Beijing, 100191, P. R. China
| | - Qingqing Li
- Department of Immunology, School of Basic Medical Sciences and NHC Key Laboratory of Medical Immunology, Peking University, Beijing, 100191, P. R. China
| | - Jia Li
- State Key Laboratory of Experimental Hematology, National Clinical Research Center for Blood Diseases, Haihe Laboratory of Cell Ecosystem, Hematopoietic Stem Cell Transplantation Center, Institute of Hematology & Blood Diseases Hospital, Chinese Academy of Medical Sciences & Peking Union Medical College, Tianjin, 300020, P. R. China
- Tianjin Institutes of Health Science, Tianjin, 301600, P. R. China
| | - Qian Ren
- State Key Laboratory of Experimental Hematology, National Clinical Research Center for Blood Diseases, Haihe Laboratory of Cell Ecosystem, Hematopoietic Stem Cell Transplantation Center, Institute of Hematology & Blood Diseases Hospital, Chinese Academy of Medical Sciences & Peking Union Medical College, Tianjin, 300020, P. R. China
- Tianjin Institutes of Health Science, Tianjin, 301600, P. R. China
| | - Runzhi Ma
- State Key Laboratory of Experimental Hematology, National Clinical Research Center for Blood Diseases, Haihe Laboratory of Cell Ecosystem, Hematopoietic Stem Cell Transplantation Center, Institute of Hematology & Blood Diseases Hospital, Chinese Academy of Medical Sciences & Peking Union Medical College, Tianjin, 300020, P. R. China
- Tianjin Institutes of Health Science, Tianjin, 301600, P. R. China
| | - Yi Zheng
- Department of Immunology, School of Basic Medical Sciences and NHC Key Laboratory of Medical Immunology, Peking University, Beijing, 100191, P. R. China
| | - Yan Zhang
- Department of Immunology, School of Basic Medical Sciences and NHC Key Laboratory of Medical Immunology, Peking University, Beijing, 100191, P. R. China
| | - Li Liu
- Tianjin First Central Hospital, Tianjin Medical University, Tianjin, 300192, P. R. China
| | - Danfeng Zheng
- Department of Immunology, School of Basic Medical Sciences and NHC Key Laboratory of Medical Immunology, Peking University, Beijing, 100191, P. R. China
| | - Pingzhang Wang
- Department of Immunology, School of Basic Medical Sciences and NHC Key Laboratory of Medical Immunology, Peking University, Beijing, 100191, P. R. China
| | - Ping Jiang
- Tianjin First Central Hospital, Tianjin Medical University, Tianjin, 300192, P. R. China
| | - Xiaoming Feng
- State Key Laboratory of Experimental Hematology, National Clinical Research Center for Blood Diseases, Haihe Laboratory of Cell Ecosystem, Hematopoietic Stem Cell Transplantation Center, Institute of Hematology & Blood Diseases Hospital, Chinese Academy of Medical Sciences & Peking Union Medical College, Tianjin, 300020, P. R. China
- Tianjin Institutes of Health Science, Tianjin, 301600, P. R. China
| | - Erlie Jiang
- State Key Laboratory of Experimental Hematology, National Clinical Research Center for Blood Diseases, Haihe Laboratory of Cell Ecosystem, Hematopoietic Stem Cell Transplantation Center, Institute of Hematology & Blood Diseases Hospital, Chinese Academy of Medical Sciences & Peking Union Medical College, Tianjin, 300020, P. R. China
- Tianjin Institutes of Health Science, Tianjin, 301600, P. R. China
| | - Ying Wang
- Department of Immunology, School of Basic Medical Sciences and NHC Key Laboratory of Medical Immunology, Peking University, Beijing, 100191, P. R. China.
| | - Sizhou Feng
- State Key Laboratory of Experimental Hematology, National Clinical Research Center for Blood Diseases, Haihe Laboratory of Cell Ecosystem, Hematopoietic Stem Cell Transplantation Center, Institute of Hematology & Blood Diseases Hospital, Chinese Academy of Medical Sciences & Peking Union Medical College, Tianjin, 300020, P. R. China.
- Tianjin Institutes of Health Science, Tianjin, 301600, P. R. China.
| |
Collapse
|
|
5
|
Lv M, Zhang J, Wang W, Jiang R, Su J. Re-identification and characterization of grass carp Ctenopharyngodon idella TLR20. FISH AND SHELLFISH IMMUNOLOGY REPORTS 2023; 5:100119. [PMID: 37841419 PMCID: PMC10568090 DOI: 10.1016/j.fsirep.2023.100119] [Citation(s) in RCA: 0] [Impact Index Per Article: 0] [Reference Citation Analysis] [Abstract] [Key Words] [Track Full Text] [Download PDF] [Figures] [Journal Information] [Subscribe] [Scholar Register] [Received: 09/10/2023] [Revised: 10/02/2023] [Accepted: 10/04/2023] [Indexed: 10/17/2023] Open
Abstract
Toll-like receptors (TLRs) play a crucial role in the recognition of microbial-associated molecular patterns in the innate immune system. Fish TLRs have undergone significant gene expansion to adapt to complex aquatic environments. Among them, TLR20 from the TLR11 family actively responds to viral and bacterial invasions. Previous studies have reported two TLR20s in grass carp (Ctenopharyngodon idella), and in this study, we revised this conclusion. Based on the latest grass carp genome, we identified a new TLR20 member. These three TLR20s are arranged in tandem on chromosome 9, indicating that they are generated by gene duplication events. They were renamed CiTLR20.1 to CiTLR20.3 based on their chromosomal positions. The CiTLR20s in C. idella exhibit higher similarities with those in Danio rerio, Cyprinus carpio, and Megalobrama amblycephala, and lower similarities with those in other distantly related fish species. Selective pressure analysis revealed low conservation and negative evolution of TLR20s during evolution. The 3D structures of the three TLR20s showed significant differences, reflecting functional variations and different downstream adaptor molecule recruitment. Transcriptome data revealed tissue distribution differences of TLR20s, with TLR20.1 showing relatively low expression levels in all the tissues, while TLR20.2 and TLR20.3 showed higher expression in the head kidney, spleen, and gill. Additionally, TLR20.2 and TLR20.3 actively responded to GCRV-II infection, with higher upregulation of TLR20.2 in response to Aeromonas hydrophila challenge. In conclusion, this study corrected the number of grass carp TLR20 members and analyzed TLR20 from an evolutionary and structural perspective, exploring its role in antiviral and antibacterial defense. This study provides reference for future research on fish TLR20.
Collapse
Affiliation(s)
- Maolin Lv
- Hubei Hongshan Laboratory, College of Fisheries, Huazhong Agricultural University, Wuhan 430070, China
- Laboratory for Marine Biology and Biotechnology, Pilot National Laboratory for Marine Science and Technology, Qingdao 266237, China
| | - Jingjing Zhang
- Hubei Hongshan Laboratory, College of Fisheries, Huazhong Agricultural University, Wuhan 430070, China
| | - Weicheng Wang
- Hubei Hongshan Laboratory, College of Fisheries, Huazhong Agricultural University, Wuhan 430070, China
| | - Rui Jiang
- Hubei Hongshan Laboratory, College of Fisheries, Huazhong Agricultural University, Wuhan 430070, China
| | - Jianguo Su
- Hubei Hongshan Laboratory, College of Fisheries, Huazhong Agricultural University, Wuhan 430070, China
- Laboratory for Marine Biology and Biotechnology, Pilot National Laboratory for Marine Science and Technology, Qingdao 266237, China
| |
Collapse
|
|
6
|
Jiang R, Zhu W, Liao Z, Yang C, Su J. TLR7 neo-functionalizes to sense dsRNA and trigger antiviral and antibacterial immunity in non-tetrapod vertebrates. iScience 2023; 26:108315. [PMID: 38025781 PMCID: PMC10679900 DOI: 10.1016/j.isci.2023.108315] [Citation(s) in RCA: 0] [Impact Index Per Article: 0] [Reference Citation Analysis] [Abstract] [Key Words] [Track Full Text] [Download PDF] [Figures] [Journal Information] [Subscribe] [Scholar Register] [Received: 07/20/2023] [Revised: 09/30/2023] [Accepted: 10/20/2023] [Indexed: 12/01/2023] Open
Abstract
TLR7 plays a crucial role in sensing viral ssRNA and initiating immune responses. Piscine TLR7 also responds to dsRNA challenge. dsRNA exists in almost all the viruses at specific stages. However, the mechanism on sensing dsRNA by TLR7 remains unknown. In the present study, we employed Ctenopharyngodon idella TLR7 (CiTLR7) to systematically explore the immune functions and mechanisms in teleost. CiTLR7 can directly bind not only ssRNA but also dsRNA at different patches in lysosome, recruit MyD88 as adaptor, and activate the downstream IFN pathway via SLC15A4/TASLa/TASLb/IRF5/IRF7 complex for antiviral and antibacterial infections and AP-1 pathway for pro-inflammatory cytokines. The key binding sites for dsRNA are L29 and L811 in CiTLR7. Further, we found that the function on recognizing dsRNA by TLR7 emerges in pisciformes and loses in tetrapods in evolution. This is the first report on sensing both ssRNA and dsRNA by a TLR member.
Collapse
Affiliation(s)
- Rui Jiang
- Hubei Hongshan Laboratory, College of Fisheries, Huazhong Agricultural University, Wuhan 430070, China
- Laboratory for Marine Biology and Biotechnology, Pilot National Laboratory for Marine Science and Technology, Qingdao 266237, China
| | - Wentao Zhu
- Hubei Hongshan Laboratory, College of Fisheries, Huazhong Agricultural University, Wuhan 430070, China
| | - Zhiwei Liao
- Hubei Hongshan Laboratory, College of Fisheries, Huazhong Agricultural University, Wuhan 430070, China
| | - Chunrong Yang
- College of Veterinary Medicine, Huazhong Agricultural University, Wuhan 430070, China
| | - Jianguo Su
- Hubei Hongshan Laboratory, College of Fisheries, Huazhong Agricultural University, Wuhan 430070, China
- Laboratory for Marine Biology and Biotechnology, Pilot National Laboratory for Marine Science and Technology, Qingdao 266237, China
| |
Collapse
|
|
7
|
Curson JE, Liu L, Luo L, Muusse TW, Lucas RM, Gunther KS, Vajjhala PR, Abrol R, Jones A, Kapetanovic R, Stacey KJ, Stow JL, Sweet MJ. TLR4 phosphorylation at tyrosine 672 activates the ERK/c-FOS signaling module for LPS-induced cytokine responses in macrophages. Eur J Immunol 2023; 53:e2250056. [PMID: 37058370 PMCID: PMC10947571 DOI: 10.1002/eji.202250056] [Citation(s) in RCA: 1] [Impact Index Per Article: 1.0] [Reference Citation Analysis] [Abstract] [Key Words] [MESH Headings] [Grants] [Track Full Text] [Journal Information] [Subscribe] [Scholar Register] [Received: 06/20/2022] [Revised: 03/20/2023] [Accepted: 04/11/2023] [Indexed: 04/15/2023]
Abstract
TLRs engage numerous adaptor proteins and signaling molecules, enabling a complex series of post-translational modifications (PTMs) to mount inflammatory responses. TLRs themselves are post-translationally modified following ligand-induced activation, with this being required to relay the full spectrum of proinflammatory signaling responses. Here, we reveal indispensable roles for TLR4 Y672 and Y749 phosphorylation in mounting optimal LPS-inducible inflammatory responses in primary mouse macrophages. LPS promotes phosphorylation at both tyrosine residues, with Y749 phosphorylation being required for maintenance of total TLR4 protein levels and Y672 phosphorylation exerting its pro-inflammatory effects more selectively by initiating ERK1/2 and c-FOS phosphorylation. Our data also support a role for the TLR4-interacting membrane proteins SCIMP and the SYK kinase axis in mediating TLR4 Y672 phosphorylation to permit downstream inflammatory responses in murine macrophages. The corresponding residue in human TLR4 (Y674) is also required for optimal LPS signaling responses. Our study, thus, reveals how a single PTM on one of the most widely studied innate immune receptors orchestrates downstream inflammatory responses.
Collapse
Affiliation(s)
- James E.B. Curson
- Institute for Molecular Bioscience (IMB)IMB Centre for Inflammation and Disease Research and Australian Infectious Diseases Research CentreThe University of QueenslandBrisbaneQueenslandAustralia
| | - Liping Liu
- Institute for Molecular Bioscience (IMB)IMB Centre for Inflammation and Disease Research and Australian Infectious Diseases Research CentreThe University of QueenslandBrisbaneQueenslandAustralia
| | - Lin Luo
- Institute for Molecular Bioscience (IMB)IMB Centre for Inflammation and Disease Research and Australian Infectious Diseases Research CentreThe University of QueenslandBrisbaneQueenslandAustralia
| | - Timothy W. Muusse
- School of Chemistry and Molecular Biosciences (SCMB) and Australian Infectious Diseases Research CentreThe University of QueenslandBrisbaneQueenslandAustralia
| | - Richard M. Lucas
- Institute for Molecular Bioscience (IMB)IMB Centre for Inflammation and Disease Research and Australian Infectious Diseases Research CentreThe University of QueenslandBrisbaneQueenslandAustralia
| | - Kimberley S. Gunther
- Institute for Molecular Bioscience (IMB)IMB Centre for Inflammation and Disease Research and Australian Infectious Diseases Research CentreThe University of QueenslandBrisbaneQueenslandAustralia
| | - Parimala R. Vajjhala
- School of Chemistry and Molecular Biosciences (SCMB) and Australian Infectious Diseases Research CentreThe University of QueenslandBrisbaneQueenslandAustralia
| | - Rishika Abrol
- Institute for Molecular Bioscience (IMB)IMB Centre for Inflammation and Disease Research and Australian Infectious Diseases Research CentreThe University of QueenslandBrisbaneQueenslandAustralia
| | - Alun Jones
- Institute for Molecular Bioscience (IMB)IMB Centre for Inflammation and Disease Research and Australian Infectious Diseases Research CentreThe University of QueenslandBrisbaneQueenslandAustralia
| | - Ronan Kapetanovic
- Institute for Molecular Bioscience (IMB)IMB Centre for Inflammation and Disease Research and Australian Infectious Diseases Research CentreThe University of QueenslandBrisbaneQueenslandAustralia
- Friedrich Miescher Institute for Biomedical ResearchBaselSwitzerland
| | - Katryn J. Stacey
- School of Chemistry and Molecular Biosciences (SCMB) and Australian Infectious Diseases Research CentreThe University of QueenslandBrisbaneQueenslandAustralia
| | - Jennifer L. Stow
- Institute for Molecular Bioscience (IMB)IMB Centre for Inflammation and Disease Research and Australian Infectious Diseases Research CentreThe University of QueenslandBrisbaneQueenslandAustralia
| | - Matthew J. Sweet
- Institute for Molecular Bioscience (IMB)IMB Centre for Inflammation and Disease Research and Australian Infectious Diseases Research CentreThe University of QueenslandBrisbaneQueenslandAustralia
| |
Collapse
|
|
8
|
Wang XY, Beeraka NM, Xue NN, Yu HM, Yang Y, Liu MX, Nikolenko VN, Liu JQ, Zhao D. Identification of a three-gene prognostic signature for radioresistant esophageal squamous cell carcinoma. World J Clin Oncol 2023; 14:13-26. [PMID: 36699628 PMCID: PMC9850665 DOI: 10.5306/wjco.v14.i1.13] [Citation(s) in RCA: 0] [Impact Index Per Article: 0] [Reference Citation Analysis] [Abstract] [Key Words] [Track Full Text] [Download PDF] [Journal Information] [Submit a Manuscript] [Subscribe] [Scholar Register] [Received: 07/25/2022] [Revised: 10/25/2022] [Accepted: 12/06/2022] [Indexed: 01/10/2023] Open
Abstract
BACKGROUND Esophageal squamous cell carcinoma (ESCC) is causing a high mortality rate due to the lack of efficient early prognosis markers and suitable therapeutic regimens. The prognostic role of genes responsible for the acquisition of radioresistance in ESCC has not been fully elucidated.
AIM To establish a prognostic model by studying gene expression patterns pertinent to radioresistance in ESCC patients.
METHODS Datasets were obtained from the Gene Expression Omnibus and The Cancer Genome Atlas databases. The edgeR, a Bioconductor package, was used to analyze mRNA expression between different groups. We screened genes specifically responsible for radioresistance to estimate overall survival. Pearson correlation analysis was performed to confirm whether the expression of those genes correlated with each other. Genes contributing to radioresistance and overall survival were assessed by the multivariate Cox regression model through the calculation of βi and risk score using the following formula: 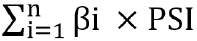 . .
RESULTS We identified three prognostic mRNAs (cathepsin S [CTSS], cluster of differentiation 180 [CD180], and SLP adapter and CSK-interacting membrane protein [SCIMP]) indicative of radioresistance. The expression of the three identified mRNAs was related to each other (r > 0.70 and P < 0.05). As to 1-year and 3-year overall survival prediction, the area under the time-dependent receiver operating characteristic curve of the signature consisting of the three mRNAs was 0.716 and 0.841, respectively. When stratifying patients based on the risk score derived from the signature, the high-risk group exhibited a higher death risk and shorter survival time than the low-risk group (P < 0.0001). Overall survival of the low-risk patients was significantly better than that of the high-risk patients (P = 0.018).
CONCLUSION We have developed a novel three-gene prognostic signature consisting of CTSS, CD180, and SCIMO for ESCC, which may facilitate the prediction of early prognosis of this malignancy.
Collapse
Affiliation(s)
- Xiao-Yan Wang
- Department of Endocrinology, The First Affiliated Hospital of Zhengzhou University, Zhengzhou 450052, Henan Province, China
| | - Narasimha M Beeraka
- Department of Radiation Oncology, The First Affiliated Hospital of Zhengzhou University, Zhengzhou 450052, Henan Province, China
- Department of Human Anatomy, I. M. Sechenov First Moscow State Medical University, Moscow 119991, Russia
- Department of Pharmaceutical Chemistry, JSS College of Pharmacy, Mysuru 570015, India
| | - Nan-Nan Xue
- Department of Radiation Oncology, The First Affiliated Hospital of Zhengzhou University, Zhengzhou 450052, Henan Province, China
| | - Hui-Ming Yu
- Department of Radiation Oncology, Peking University Cancer Hospital & Institute, Beijing 065005, China
| | - Ya Yang
- Department of Radiation Oncology, The First Affiliated Hospital of Zhengzhou University, Zhengzhou 450052, Henan Province, China
| | - Mao-Xing Liu
- Key Laboratory of Carcinogenesis and Translational Research (Ministry of Education), Department of Gastrointestinal Surgery IV, Peking University Cancer Hospital & Institute, Beijing, China
| | - Vladimir N Nikolenko
- Department of Human Anatomy, I. M. Sechenov First Moscow State Medical University, Moscow 119991, Russia
- M.V. Lomonosov Moscow State University, Moscow 119991, Russia
| | - Jun-Qi Liu
- Department of Radiation Oncology, The First Affiliated Hospital of Zhengzhou University, Zhengzhou 450052, Henan Province, China
| | - Di Zhao
- Department of Endocrinology, The First Affiliated Hospital of Zhengzhou University, Zhengzhou 450052, Henan Province, China
| |
Collapse
|
|
9
|
Metabolism-dependent ferroptosis promotes mitochondrial dysfunction and inflammation in CD4 + T lymphocytes in HIV-infected immune non-responders. EBioMedicine 2022; 86:104382. [PMID: 36462403 PMCID: PMC9718960 DOI: 10.1016/j.ebiom.2022.104382] [Citation(s) in RCA: 15] [Impact Index Per Article: 7.5] [Reference Citation Analysis] [Abstract] [Key Words] [Track Full Text] [Download PDF] [Figures] [Journal Information] [Subscribe] [Scholar Register] [Received: 07/07/2022] [Revised: 10/29/2022] [Accepted: 11/09/2022] [Indexed: 12/03/2022] Open
Abstract
BACKGROUND HIV immune non-responders (INRs) are described as a failure to reestablish a pool of CD4+ T lymphocytes (CD4 cells) after antiretroviral therapy (ART), which is related to poor clinical results. Ferroptosis is a newly discovered form of cell death characterised by iron-dependent lipid peroxidation and the accumulation of reactive oxygen species (ROS). The mechanism of unrecoverable CD4 cells in INRs and whether ferroptosis plays a role are not fully understood. METHODS Ninety-two people living with HIV (PLHIVs) who experienced four-year ART with sustained viral suppression, including 27 INRs, 34 partial responders (PRs), and 31 complete responders (CRs); and 26 uninfected control participants (UCs) were analysed for 16 immune parameters with flow cytometry. Then plasma lipid, iron and oxidation, and antioxidant indicators were detected by ELISA, and CD4 cells were sorted out and visualised under transmission electron microscopy. Finally, ferroptosis inhibitors were added, and alterations in CD4 cell phenotype and function were observed. FINDINGS We found decreased recent thymic emigrants (RTE), over-activation and over-proliferation phenotypes, diminished killing function, decreased IL-7R and more severe inflammation; increased lipid peroxidation in the mitochondria and disruptions of the mitochondrial structure, showing typical features of ferroptosis in CD4 cells in INRs. Additionally, ferroptosis inhibitors could reduce inflammation and repair mitochondrial damage. Meanwhile, ELISA results showed increased plasma free fatty acids (FFA) and an imbalance of oxidative and antioxidant systems in INRs. Flow cytometry results displayed alterations of both transferrin receptor (CD71) and lipid transporter (CD36) expressions on the surface of CD4 cells. Mechanistically, there was a stronger correlation between CD36 expression and mitochondrial lipid peroxidation production, ferroptosis makers, and inflammation indicators; while amino acid transporter (CD98) was more related to killing functions; and CD71 was more closely related to activation status in CD4 cells. INTERPRETATION Cellular metabolism was closely correlated with its diverse functions in INRs. Ferroptosis was observed in CD4 cells of INRs, and inhibiting ferroptosis through modulating mitochondrial disorders and inflammation may offer an alternative immunological strategy for reinvigorating CD4 cells in INRs. FUNDING This research was supported by the 13th Five-year Plan, Ministry of Science and Technology of China (2018ZX10302-102), Beijing Municipal Administration of Hospitals' Ascent Plan (DFL20191802), and Beijing Municipal Administration of Hospitals Clinical Medicine Development of Special Funding Support (ZYLX202126).
Collapse
|
|
10
|
Lopes N, Galluso J, Escalière B, Carpentier S, Kerdiles YM, Vivier E. Tissue-specific transcriptional profiles and heterogeneity of natural killer cells and group 1 innate lymphoid cells. Cell Rep Med 2022; 3:100812. [PMID: 36384102 PMCID: PMC9729827 DOI: 10.1016/j.xcrm.2022.100812] [Citation(s) in RCA: 12] [Impact Index Per Article: 6.0] [Reference Citation Analysis] [Abstract] [Key Words] [Track Full Text] [Download PDF] [Figures] [Journal Information] [Subscribe] [Scholar Register] [Received: 05/31/2022] [Revised: 08/18/2022] [Accepted: 10/17/2022] [Indexed: 11/17/2022]
Abstract
Natural killer (NK) cells and type 1 innate lymphoid cells (ILC1s) are populations of non-T, non-B lymphocytes in peripheral tissues. Although NK and ILC1 subsets have been described, their identification and characteristics remain unclear. We performed single-cell RNA sequencing and CITE-seq to explore NK and ILC1 heterogeneity between tissues. We observed that although NK1 and NK2 subsets are conserved in spleen and liver, ILC1s are heterogeneous across tissues. We identified sets of genes expressed by related subsets or characterizing unique ILC1 populations in each organ. The syndecan-4 appeared as a marker discriminating murine ILC1 from NK cells across organs. Finally, we revealed that the expressions of EOMES, GZMA, IRF8, JAK1, NKG7, PLEK, PRF1, and ZEB2 define NK cells and that IL7R, LTB, and RGS1 differentiate ILC1s from NK cells in mice and humans. Our data constitute an important resource to improve our understanding of NK-ILC1 origin, phenotype, and biology.
Collapse
Affiliation(s)
- Noella Lopes
- Aix Marseille Université, CNRS, INSERM, Centre d'Immunologie de Marseille-Luminy, Marseille, France
| | - Justine Galluso
- Aix Marseille Université, CNRS, INSERM, Centre d'Immunologie de Marseille-Luminy, Marseille, France
| | - Bertrand Escalière
- Aix Marseille Université, CNRS, INSERM, Centre d'Immunologie de Marseille-Luminy, Marseille, France
| | | | - Yann M. Kerdiles
- Aix Marseille Université, CNRS, INSERM, Centre d'Immunologie de Marseille-Luminy, Marseille, France,Corresponding author
| | - Eric Vivier
- Aix Marseille Université, CNRS, INSERM, Centre d'Immunologie de Marseille-Luminy, Marseille, France,Innate Pharma Research Laboratories, Innate Pharma, Marseille, France,APHM, Hôpital de la Timone, Marseille-Immunopôle, Marseille, France,Corresponding author
| |
Collapse
|
|
11
|
Luan C, Jin S, Hu Y, Zhou X, Liu L, Li R, Ju M, Huang D, Chen K. Whole-genome identification and construction of the lncRNA-mRNA co-expression network in patients with actinic keratosis. Transl Cancer Res 2022; 11:4070-4078. [PMID: 36523309 PMCID: PMC9745357 DOI: 10.21037/tcr-22-842] [Citation(s) in RCA: 0] [Impact Index Per Article: 0] [Reference Citation Analysis] [Abstract] [Key Words] [Track Full Text] [Journal Information] [Subscribe] [Scholar Register] [Received: 03/28/2022] [Accepted: 08/17/2022] [Indexed: 08/30/2023]
Abstract
BACKGROUND Actinic keratosis (AK) is a common premalignant lesion induced by chronic exposure to ultraviolet radiation and may develop into invasive cutaneous squamous carcinoma (cSCC). The identification of specific biomarkers in AK are still unclear. Long non-coding RNAs (lncRNAs), as transcripts of more than 200 nucleotides, significantly involving in multiple biologic processes, especially in the development of tumors. METHODS In our study, we obtained data from RNA-sequencing analysis using two AK lesion tissues and three normal cutaneous tissues to comparatively analyze the differentially expressed (DE) lncRNAs and messenger RNAs (mRNAs). Firstly, we used microarray analyses to identify DE lncRNAs and DE mRNAs. Secondly, we performed Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis to analyze the primary function and find out significant pathways of these DE mRNA and lncRNAs. Finally, we used the top ten DE lncRNAs to construct a lncRNA-mRNA co-expression network. RESULTS Our results showed that there were a total of 2,097 DE lncRNAs and 2,043 DE mRNAs identified. GO and KEGG analysis and the lncRNA-mRNA co-expression network (using the top 10 DE lncRNAs comprises 130 specific co-expressed mRNAs to construct) indicated that lncRNA uc011fnr.2 may negatively regulate SCIMP and Toll-like receptor 4 (TLR4) and play an important role in Janus kinase-signal transducer and activator of transcription 3 (JAK-STAT3) signaling pathway of AK. CONCLUSIONS lncRNA uc011fnr.2 may play an important role in JAK-STAT3 signaling pathway of AK by modulating SCIMP, TLR4 and IL-6. Further research is required to validate the value of lncRNA uc011fnr.2 in the progression of AK.
Collapse
Affiliation(s)
- Chao Luan
- Institute of Dermatology, Jiangsu Key Laboratory of Molecular Biology for Skin Diseases and STIs, Chinese Academy of Medical Science & Peking Union Medical College, Nanjing, China
| | - Shuang Jin
- Institute of Dermatology, Jiangsu Key Laboratory of Molecular Biology for Skin Diseases and STIs, Chinese Academy of Medical Science & Peking Union Medical College, Nanjing, China
| | - Yu Hu
- Institute of Dermatology, Jiangsu Key Laboratory of Molecular Biology for Skin Diseases and STIs, Chinese Academy of Medical Science & Peking Union Medical College, Nanjing, China
| | - Xuyue Zhou
- Institute of Dermatology, Jiangsu Key Laboratory of Molecular Biology for Skin Diseases and STIs, Chinese Academy of Medical Science & Peking Union Medical College, Nanjing, China
| | - Lingxi Liu
- Institute of Dermatology, Jiangsu Key Laboratory of Molecular Biology for Skin Diseases and STIs, Chinese Academy of Medical Science & Peking Union Medical College, Nanjing, China
| | - Rong Li
- Institute of Dermatology, Jiangsu Key Laboratory of Molecular Biology for Skin Diseases and STIs, Chinese Academy of Medical Science & Peking Union Medical College, Nanjing, China
| | - Mei Ju
- Institute of Dermatology, Jiangsu Key Laboratory of Molecular Biology for Skin Diseases and STIs, Chinese Academy of Medical Science & Peking Union Medical College, Nanjing, China
| | - Dan Huang
- Institute of Dermatology, Jiangsu Key Laboratory of Molecular Biology for Skin Diseases and STIs, Chinese Academy of Medical Science & Peking Union Medical College, Nanjing, China
| | - Kun Chen
- Institute of Dermatology, Jiangsu Key Laboratory of Molecular Biology for Skin Diseases and STIs, Chinese Academy of Medical Science & Peking Union Medical College, Nanjing, China
| |
Collapse
|
|
12
|
Lucas RM, Luo L, Stow JL. ERK1/2 in immune signalling. Biochem Soc Trans 2022; 50:1341-1352. [PMID: 36281999 PMCID: PMC9704528 DOI: 10.1042/bst20220271] [Citation(s) in RCA: 25] [Impact Index Per Article: 12.5] [Reference Citation Analysis] [Abstract] [Key Words] [MESH Headings] [Track Full Text] [Journal Information] [Subscribe] [Scholar Register] [Received: 06/12/2022] [Revised: 10/07/2022] [Accepted: 10/10/2022] [Indexed: 07/30/2023]
Abstract
Extracellular signal-related kinases 1 and 2 (ERK1/2) are the final components of the mitogen-activated protein kinase (MAPK) phosphorylation cascade, an integral module in a diverse array of signalling pathways for shaping cell behaviour and fate. More recently, studies have shown that ERK1/2 plays an essential role downstream of immune receptors to elicit inflammatory gene expression in response to infection and cell or tissue damage. Much of this work has studied ERK1/2 activation in Toll-like receptor (TLR) pathways, providing mechanistic insights into its recruitment, compartmentalisation and activation in cells of the innate immune system. In this review, we summarise the typical activation of ERK1/2 in growth factor receptor pathways before discussing its known roles in immune cell signalling with a focus downstream of TLRs. We examine emerging research uncovering evidence of dysfunctional ERK1/2 signalling in inflammatory diseases and discuss the potential therapeutic benefit of targeting ERK1/2 pathways in inflammation.
Collapse
Affiliation(s)
- Richard M. Lucas
- Institute for Molecular Bioscience (IMB) and Centre for Inflammation and Disease Research, The University of Queensland, St Lucia, QLD 4072, Australia
| | - Lin Luo
- Institute for Molecular Bioscience (IMB) and Centre for Inflammation and Disease Research, The University of Queensland, St Lucia, QLD 4072, Australia
| | - Jennifer L. Stow
- Institute for Molecular Bioscience (IMB) and Centre for Inflammation and Disease Research, The University of Queensland, St Lucia, QLD 4072, Australia
| |
Collapse
|
|
13
|
The transmembrane adapter SCIMP recruits tyrosine kinase Syk to phosphorylate Toll-like receptors to mediate selective inflammatory outputs. J Biol Chem 2022; 298:101857. [PMID: 35337798 PMCID: PMC9052152 DOI: 10.1016/j.jbc.2022.101857] [Citation(s) in RCA: 5] [Impact Index Per Article: 2.5] [Reference Citation Analysis] [Abstract] [Key Words] [Track Full Text] [Download PDF] [Figures] [Journal Information] [Subscribe] [Scholar Register] [Received: 11/07/2021] [Revised: 03/08/2022] [Accepted: 03/11/2022] [Indexed: 11/23/2022] Open
Abstract
Innate immune signaling by Toll-like receptors (TLRs) involves receptor phosphorylation, which helps to shape and drive key inflammatory outputs, yet our understanding of the kinases and mechanisms that mediate TLR phosphorylation is incomplete. Spleen tyrosine kinase (Syk) is a nonreceptor protein tyrosine kinase, which is known to relay adaptive and innate immune signaling, including from TLRs. However, TLRs do not contain the conserved dual immunoreceptor tyrosine-based activation motifs that typically recruit Syk to many other receptors. One possibility is that the Syk-TLR association is indirect, relying on an intermediary scaffolding protein. We previously identified a role for the palmitoylated transmembrane adapter protein SCIMP in scaffolding the Src tyrosine kinase Lyn, for TLR phosphorylation, but the role of SCIMP in mediating the interaction between Syk and TLRs has not yet been investigated. Here, we show that SCIMP recruits Syk in response to lipopolysaccharide-mediated TLR4 activation. We also show that Syk contributes to the phosphorylation of SCIMP and TLR4 to enhance their binding. Further evidence pinpoints two specific phosphorylation sites in SCIMP critical for its interaction with Syk-SH2 domains in the absence of immunoreceptor tyrosine-based activation motifs. Finally, using inhibitors and primary macrophages from SCIMP-/- mice, we confirm a functional role for SCIMP-mediated Syk interaction in modulating TLR4 phosphorylation, signaling, and cytokine outputs. In conclusion, we identify SCIMP as a novel, immune-specific Syk scaffold, which can contribute to inflammation through selective TLR-driven inflammatory responses.
Collapse
|
|
14
|
Curson JE, Luo L, Liu L, Burgess BJ, Bokil NJ, Wall AA, Brdicka T, Kapetanovic R, Stow JL, Sweet MJ. An alternative downstream translation start site in the non-TIR adaptor Scimp enables selective amplification of CpG DNA responses in mouse macrophages. Immunol Cell Biol 2022; 100:267-284. [PMID: 35201640 PMCID: PMC9544816 DOI: 10.1111/imcb.12540] [Citation(s) in RCA: 4] [Impact Index Per Article: 2.0] [Reference Citation Analysis] [Abstract] [Key Words] [Track Full Text] [Download PDF] [Figures] [Journal Information] [Subscribe] [Scholar Register] [Received: 09/03/2021] [Revised: 02/17/2022] [Accepted: 02/22/2022] [Indexed: 01/01/2023]
Abstract
Toll-like receptor (TLR) signaling relies on Toll/interleukin-1 receptor homology (TIR) domain-containing adaptor proteins that recruit downstream signaling molecules to generate tailored immune responses. In addition, the palmitoylated transmembrane adaptor protein family member Scimp acts as a non-TIR-containing adaptor protein in macrophages, scaffolding the Src family kinase Lyn to enable TLR phosphorylation and proinflammatory signaling responses. Here we report the existence of a smaller, naturally occurring translational variant of Scimp (Scimp TV1), which is generated through leaky scanning and translation at a downstream methionine. Scimp TV1 also scaffolds Lyn, but in contrast to full-length Scimp, it is basally rather than lipopolysaccharide (LPS)-inducibly phosphorylated. Macrophages from mice that selectively express Scimp TV1, but not full-length Scimp, have impaired sustained LPS-inducible cytokine responses. Furthermore, in granulocyte macrophage colony-stimulating factor-derived myeloid cells that express high levels of Scimp, selective overexpression of Scimp TV1 enhances CpG DNA-inducible cytokine production. Unlike full-length Scimp that localizes to the cell surface and filopodia, Scimp TV1 accumulates in intracellular compartments, particularly the Golgi. Moreover, this variant of Scimp is not inducibly phosphorylated in response to CpG DNA, suggesting that it may act via an indirect mechanism to enhance TLR9 responses. Our findings thus reveal the use of alternative translation start sites as a previously unrecognized mechanism for diversifying TLR responses in the innate immune system.
Collapse
Affiliation(s)
- James Eb Curson
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research, and Australian Infectious Diseases Research Centre, The University of Queensland, Brisbane, QLD, Australia
| | - Lin Luo
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research, and Australian Infectious Diseases Research Centre, The University of Queensland, Brisbane, QLD, Australia
| | - Liping Liu
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research, and Australian Infectious Diseases Research Centre, The University of Queensland, Brisbane, QLD, Australia
| | - Belinda J Burgess
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research, and Australian Infectious Diseases Research Centre, The University of Queensland, Brisbane, QLD, Australia
| | - Nilesh J Bokil
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research, and Australian Infectious Diseases Research Centre, The University of Queensland, Brisbane, QLD, Australia
| | - Adam A Wall
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research, and Australian Infectious Diseases Research Centre, The University of Queensland, Brisbane, QLD, Australia
| | - Tomas Brdicka
- Laboratory of Leukocyte Signaling, Institute of Molecular Genetics of the Czech Academy of Sciences, Prague, Czech Republic
| | - Ronan Kapetanovic
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research, and Australian Infectious Diseases Research Centre, The University of Queensland, Brisbane, QLD, Australia.,Friedrich Miescher Institute for Biomedical Research, Basel, Switzerland
| | - Jennifer L Stow
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research, and Australian Infectious Diseases Research Centre, The University of Queensland, Brisbane, QLD, Australia
| | - Matthew J Sweet
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research, and Australian Infectious Diseases Research Centre, The University of Queensland, Brisbane, QLD, Australia
| |
Collapse
|
|
15
|
Yang JX, Tseng JC, Yu GY, Luo Y, Huang CYF, Hong YR, Chuang TH. Recent Advances in the Development of Toll-like Receptor Agonist-Based Vaccine Adjuvants for Infectious Diseases. Pharmaceutics 2022; 14:pharmaceutics14020423. [PMID: 35214155 PMCID: PMC8878135 DOI: 10.3390/pharmaceutics14020423] [Citation(s) in RCA: 37] [Impact Index Per Article: 18.5] [Reference Citation Analysis] [Abstract] [Track Full Text] [Download PDF] [Figures] [Journal Information] [Subscribe] [Scholar Register] [Received: 01/13/2022] [Revised: 02/11/2022] [Accepted: 02/14/2022] [Indexed: 02/06/2023] Open
Abstract
Vaccines are powerful tools for controlling microbial infections and preventing epidemic diseases. Efficient inactive, subunit, or viral-like particle vaccines usually rely on a safe and potent adjuvant to boost the immune response to the antigen. After a slow start, over the last decade there has been increased developments on adjuvants for human vaccines. The development of adjuvants has paralleled our increased understanding of the molecular mechanisms for the pattern recognition receptor (PRR)-mediated activation of immune responses. Toll-like receptors (TLRs) are a group of PRRs that recognize microbial pathogens to initiate a host’s response to infection. Activation of TLRs triggers potent and immediate innate immune responses, which leads to subsequent adaptive immune responses. Therefore, these TLRs are ideal targets for the development of effective adjuvants. To date, TLR agonists such as monophosphoryl lipid A (MPL) and CpG-1018 have been formulated in licensed vaccines for their adjuvant activity, and other TLR agonists are being developed for this purpose. The COVID-19 pandemic has also accelerated clinical research of vaccines containing TLR agonist-based adjuvants. In this paper, we reviewed the agonists for TLR activation and the molecular mechanisms associated with the adjuvants’ effects on TLR activation, emphasizing recent advances in the development of TLR agonist-based vaccine adjuvants for infectious diseases.
Collapse
Affiliation(s)
- Jing-Xing Yang
- Immunology Research Center, National Health Research Institutes, Miaoli 35053, Taiwan; (J.-X.Y.); (J.-C.T.)
| | - Jen-Chih Tseng
- Immunology Research Center, National Health Research Institutes, Miaoli 35053, Taiwan; (J.-X.Y.); (J.-C.T.)
| | - Guann-Yi Yu
- National Institute of Infectious Diseases and Vaccinology, National Health Research Institutes, Miaoli 35053, Taiwan;
| | - Yunping Luo
- Department of Immunology, Institute of Basic Medical Sciences, Chinese Academy of Medical Sciences, School of Basic Medicine, Peking Union Medical College, Beijing 100005, China;
| | - Chi-Ying F. Huang
- Institute of Biopharmaceutical Sciences, College of Pharmaceutical Sciences, National Yang Ming Chiao Tung University, Taipei 112304, Taiwan;
| | - Yi-Ren Hong
- Graduate Institute of Medicine, College of Medicine, Kaohsiung Medical University, Kaohsiung 80708, Taiwan;
| | - Tsung-Hsien Chuang
- Immunology Research Center, National Health Research Institutes, Miaoli 35053, Taiwan; (J.-X.Y.); (J.-C.T.)
- Department of Life Sciences, National Central University, Taoyuan City 32001, Taiwan
- Program in Environmental and Occupational Medicine, Kaohsiung Medical University, Kaohsiung 80708, Taiwan
- Correspondence: ; Tel.: +886-37-246166 (ext. 37611)
| |
Collapse
|
|
16
|
Souza SP, Splitt SD, Sànchez-Arcila JC, Alvarez JA, Wilson JN, Wizzard S, Luo Z, Baumgarth N, Jensen KDC. Genetic mapping reveals Nfkbid as a central regulator of humoral immunity to Toxoplasma gondii. PLoS Pathog 2021; 17:e1010081. [PMID: 34871323 PMCID: PMC8675933 DOI: 10.1371/journal.ppat.1010081] [Citation(s) in RCA: 7] [Impact Index Per Article: 2.3] [Reference Citation Analysis] [Abstract] [MESH Headings] [Grants] [Track Full Text] [Download PDF] [Figures] [Journal Information] [Subscribe] [Scholar Register] [Received: 05/20/2021] [Revised: 12/16/2021] [Accepted: 11/01/2021] [Indexed: 12/29/2022] Open
Abstract
Protective immunity to parasitic infections has been difficult to elicit by vaccines. Among parasites that evade vaccine-induced immunity is Toxoplasma gondii, which causes lethal secondary infections in chronically infected mice. Here we report that unlike susceptible C57BL/6J mice, A/J mice were highly resistant to secondary infection. To identify correlates of immunity, we utilized forward genetics to identify Nfkbid, a nuclear regulator of NF-κB that is required for B cell activation and B-1 cell development. Nfkbid-null mice (“bumble”) did not generate parasite-specific IgM and lacked robust parasite-specific IgG, which correlated with defects in B-2 cell maturation and class-switch recombination. Though high-affinity antibodies were B-2 derived, transfer of B-1 cells partially rescued the immunity defects observed in bumble mice and were required for 100% vaccine efficacy in bone marrow chimeric mice. Immunity in resistant mice correlated with robust isotype class-switching in both B cell lineages, which can be fine-tuned by Nfkbid gene expression. We propose a model whereby humoral immunity to T. gondii is regulated by Nfkbid and requires B-1 and B-2 cells for full protection. Eukaryotic parasitic diseases account for approximately one fifth of all childhood deaths, yet no highly protective vaccine exists for any human parasite. More research must be done to discover how to elicit protective vaccine-induced immunity to parasitic pathogens. We used an unbiased genetic screen to find key genes responsible for immunity to the eukaryotic parasite Toxoplasma gondii. Our screen found Nfkbid, a transcription factor regulator, which controls B cell activation and innate-like B-1 cell development. Mice without Nfkbid were not protected against T. gondii and were deficient at making antibodies against the parasite. Our survival studies of vaccinated mice with and without B-1 compartments found that B-1 cells improved survival, suggesting that B-1 cells act in conjunction with B-2 cells to provide vaccine-induced immunity. Nfkbid and other loci identified in our unbiased screen represent potential targets for vaccines to elicit protective immune responses against parasitic pathogens.
Collapse
Affiliation(s)
- Scott P. Souza
- School of Natural Sciences, Department of Molecular and Cell Biology, University of California, Merced, Merced, California, United States of America
- Graduate Program in Quantitative and Systems Biology, University of California, Merced, Merced, California, United States of America
| | - Samantha D. Splitt
- School of Natural Sciences, Department of Molecular and Cell Biology, University of California, Merced, Merced, California, United States of America
- Graduate Program in Quantitative and Systems Biology, University of California, Merced, Merced, California, United States of America
| | - Juan C. Sànchez-Arcila
- School of Natural Sciences, Department of Molecular and Cell Biology, University of California, Merced, Merced, California, United States of America
| | - Julia A. Alvarez
- School of Natural Sciences, Department of Molecular and Cell Biology, University of California, Merced, Merced, California, United States of America
- Graduate Program in Quantitative and Systems Biology, University of California, Merced, Merced, California, United States of America
| | - Jessica N. Wilson
- School of Natural Sciences, Department of Molecular and Cell Biology, University of California, Merced, Merced, California, United States of America
- Graduate Program in Quantitative and Systems Biology, University of California, Merced, Merced, California, United States of America
| | - Safuwra Wizzard
- School of Natural Sciences, Department of Molecular and Cell Biology, University of California, Merced, Merced, California, United States of America
| | - Zheng Luo
- Center for Immunology & Infectious Diseases, and Department of Pathology, Microbiology and Immunology, University of California, Davis, Davis, California, United States of America
| | - Nicole Baumgarth
- Center for Immunology & Infectious Diseases, and Department of Pathology, Microbiology and Immunology, University of California, Davis, Davis, California, United States of America
| | - Kirk D. C. Jensen
- School of Natural Sciences, Department of Molecular and Cell Biology, University of California, Merced, Merced, California, United States of America
- Health Science Research Institute, University of California, Merced, Merced, California, United States of America
- * E-mail:
| |
Collapse
|
|
17
|
Li Y, Laws SM, Miles LA, Wiley JS, Huang X, Masters CL, Gu BJ. Genomics of Alzheimer's disease implicates the innate and adaptive immune systems. Cell Mol Life Sci 2021; 78:7397-7426. [PMID: 34708251 PMCID: PMC11073066 DOI: 10.1007/s00018-021-03986-5] [Citation(s) in RCA: 26] [Impact Index Per Article: 8.7] [Reference Citation Analysis] [Abstract] [Key Words] [MESH Headings] [Grants] [Track Full Text] [Journal Information] [Subscribe] [Scholar Register] [Received: 06/16/2021] [Revised: 09/14/2021] [Accepted: 10/16/2021] [Indexed: 02/08/2023]
Abstract
Alzheimer's disease (AD) is a chronic neurodegenerative disease characterised by cognitive impairment, behavioural alteration, and functional decline. Over 130 AD-associated susceptibility loci have been identified by genome-wide association studies (GWAS), while whole genome sequencing (WGS) and whole exome sequencing (WES) studies have identified AD-associated rare variants. These variants are enriched in APOE, TREM2, CR1, CD33, CLU, BIN1, CD2AP, PILRA, SCIMP, PICALM, SORL1, SPI1, RIN3, and more genes. Given that aging is the single largest risk factor for late-onset AD (LOAD), the accumulation of somatic mutations in the brain and blood of AD patients have also been explored. Collectively, these genetic findings implicate the role of innate and adaptive immunity in LOAD pathogenesis and suggest that a systemic failure of cell-mediated amyloid-β (Aβ) clearance contributes to AD onset and progression. AD-associated variants are particularly enriched in myeloid-specific regulatory regions, implying that AD risk variants are likely to perturbate the expression of myeloid-specific AD-associated genes to interfere Aβ clearance. Defective phagocytosis, endocytosis, and autophagy may drive Aβ accumulation, which may be related to naturally-occurring antibodies to Aβ (Nabs-Aβ) produced by adaptive responses. Passive immunisation is providing efficiency in clearing Aβ and slowing cognitive decline, such as aducanumab, donanemab, and lecanemab (ban2401). Causation of AD by impairment of the innate immunity and treatment using the tools of adaptive immunity is emerging as a new paradigm for AD, but immunotherapy that boosts the innate immune functions of myeloid cells is highly expected to modulate disease progression at asymptomatic stage.
Collapse
Affiliation(s)
- Yihan Li
- The Florey Institute, The University of Melbourne, 30 Royal Parade, Parkville, VIC, 3052, Australia
| | - Simon M Laws
- Centre for Precision Health, Edith Cowan University, 270 Joondalup Dr, Joondalup, WA, 6027, Australia
- Collaborative Genomics and Translation Group, School of Medical and Health Sciences, Edith Cowan University, 270 Joondalup Dr, Joondalup, WA, 6027, Australia
| | - Luke A Miles
- The Florey Institute, The University of Melbourne, 30 Royal Parade, Parkville, VIC, 3052, Australia
| | - James S Wiley
- The Florey Institute, The University of Melbourne, 30 Royal Parade, Parkville, VIC, 3052, Australia
| | - Xin Huang
- The Florey Institute, The University of Melbourne, 30 Royal Parade, Parkville, VIC, 3052, Australia
| | - Colin L Masters
- The Florey Institute, The University of Melbourne, 30 Royal Parade, Parkville, VIC, 3052, Australia
| | - Ben J Gu
- The Florey Institute, The University of Melbourne, 30 Royal Parade, Parkville, VIC, 3052, Australia.
| |
Collapse
|
|
18
|
Ramnath D, Das Gupta K, Wang Y, Abrol R, Curson JEB, Lim J, Reid RC, Mansell A, Blumenthal A, Karunakaran D, Fairlie DP, Sweet MJ. The histone deacetylase Hdac7 supports LPS-inducible glycolysis and Il-1β production in murine macrophages via distinct mechanisms. J Leukoc Biol 2021; 111:327-336. [PMID: 34811804 DOI: 10.1002/jlb.2mr1021-260r] [Citation(s) in RCA: 2] [Impact Index Per Article: 0.7] [Reference Citation Analysis] [Abstract] [Key Words] [Track Full Text] [Journal Information] [Subscribe] [Scholar Register] [Received: 06/13/2021] [Revised: 10/05/2021] [Accepted: 11/03/2021] [Indexed: 12/18/2022] Open
Abstract
TLRs reprogram macrophage metabolism, enhancing glycolysis and promoting flux through the tricarboxylic acid cycle to enable histone acetylation and inflammatory gene expression. The histone deacetylase (HDAC) family of lysine deacetylases regulates both TLR-inducible glycolysis and inflammatory responses. Here, we show that the TLR4 agonist LPS, as well as agonists of other TLRs, rapidly increase enzymatic activity of the class IIa HDAC family (HDAC4, 5, 7, 9) in both primary human and murine macrophages. This response was abrogated in murine macrophages deficient in histone deacetylase 7 (Hdac7), highlighting a selective role for this specific lysine deacetylase during immediate macrophage activation. With the exception of the TLR3 agonist polyI:C, TLR-inducible activation of Hdac7 enzymatic activity required the MyD88 adaptor protein. The rapid glycolysis response, as assessed by extracellular acidification rate, was attenuated in Hdac7-deficient mouse macrophages responding to submaximal LPS concentrations. Surprisingly however, reconstitution of these cells with either wild-type or an enzyme-dead mutant of Hdac7 enhanced LPS-inducible glycolysis, whereas only the former promoted production of the inflammatory mediators Il-1β and Ccl2. Thus, Hdac7 enzymatic activity is required for TLR-inducible production of specific inflammatory mediators, whereas it acts in an enzyme-independent fashion to reprogram metabolism in macrophages responding to submaximal LPS concentrations. Hdac7 is thus a bifurcation point for regulated metabolism and inflammatory responses in macrophages. Taken together with existing literature, our findings support a model in which submaximal and maximal activation of macrophages via TLR4 instruct glycolysis through distinct mechanisms, leading to divergent biological responses.
Collapse
Affiliation(s)
- Divya Ramnath
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, Queensland, Australia.,Australian Infectious Diseases Research Centre, The University of Queensland, Brisbane, Queensland, Australia
| | - Kaustav Das Gupta
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, Queensland, Australia.,Australian Infectious Diseases Research Centre, The University of Queensland, Brisbane, Queensland, Australia
| | - Yizhuo Wang
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, Queensland, Australia.,Australian Infectious Diseases Research Centre, The University of Queensland, Brisbane, Queensland, Australia
| | - Rishika Abrol
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, Queensland, Australia.,Australian Infectious Diseases Research Centre, The University of Queensland, Brisbane, Queensland, Australia
| | - James E B Curson
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, Queensland, Australia.,Australian Infectious Diseases Research Centre, The University of Queensland, Brisbane, Queensland, Australia
| | - Junxian Lim
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, Queensland, Australia.,Australian Research Council Centre of Excellence in Advanced Molecular Imaging, The University of Queensland, Brisbane, Queensland, Australia
| | - Robert C Reid
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, Queensland, Australia.,Australian Research Council Centre of Excellence in Advanced Molecular Imaging, The University of Queensland, Brisbane, Queensland, Australia
| | - Ashley Mansell
- Hudson Institute of Medical Research, Clayton, Victoria, Australia
| | - Antje Blumenthal
- The University of Queensland Diamantina Institute, The University of Queensland, Brisbane, Queensland, Australia
| | - Denuja Karunakaran
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, Queensland, Australia.,Australian Infectious Diseases Research Centre, The University of Queensland, Brisbane, Queensland, Australia
| | - David P Fairlie
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, Queensland, Australia.,Australian Infectious Diseases Research Centre, The University of Queensland, Brisbane, Queensland, Australia.,Australian Research Council Centre of Excellence in Advanced Molecular Imaging, The University of Queensland, Brisbane, Queensland, Australia
| | - Matthew J Sweet
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, Queensland, Australia.,Australian Infectious Diseases Research Centre, The University of Queensland, Brisbane, Queensland, Australia
| |
Collapse
|
|
19
|
Lucas RM, Liu L, Curson JEB, Koh YWH, Tuladhar N, Condon ND, Das Gupta K, Burgener SS, Schroder K, Ingley E, Sweet MJ, Stow JL, Luo L. SCIMP is a spatiotemporal transmembrane scaffold for Erk1/2 to direct pro-inflammatory signaling in TLR-activated macrophages. Cell Rep 2021; 36:109662. [PMID: 34496234 DOI: 10.1016/j.celrep.2021.109662] [Citation(s) in RCA: 9] [Impact Index Per Article: 3.0] [Reference Citation Analysis] [Abstract] [Key Words] [Track Full Text] [Journal Information] [Subscribe] [Scholar Register] [Received: 03/15/2021] [Revised: 07/12/2021] [Accepted: 08/13/2021] [Indexed: 02/06/2023] Open
Abstract
Immune cells are armed with Toll-like receptors (TLRs) for sensing and responding to pathogens and other danger cues. The role of extracellular-signal-regulated kinases 1/2 (Erk1/2) in TLR signaling remains enigmatic, with both pro- and anti-inflammatory functions described. We reveal here that the immune-specific transmembrane adaptor SCIMP is a direct scaffold for Erk1/2 in TLR pathways, with high-resolution, live-cell imaging revealing that SCIMP guides the spatial and temporal recruitment of Erk2 to membrane ruffles and macropinosomes for pro-inflammatory TLR4 signaling. SCIMP-deficient mice display defects in Erk1/2 recruitment to TLR4, c-Fos activation, and pro-inflammatory cytokine production, with these effects being phenocopied by Erk1/2 signaling inhibition. Our findings thus delineate a selective role for SCIMP as a key scaffold for the membrane recruitment of Erk1/2 kinase to initiate TLR-mediated pro-inflammatory responses in macrophages.
Collapse
Affiliation(s)
- Richard M Lucas
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, QLD 4072, Australia
| | - Liping Liu
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, QLD 4072, Australia
| | - James E B Curson
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, QLD 4072, Australia
| | - Yvette W H Koh
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, QLD 4072, Australia
| | - Neeraj Tuladhar
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, QLD 4072, Australia
| | - Nicholas D Condon
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, QLD 4072, Australia
| | - Kaustav Das Gupta
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, QLD 4072, Australia
| | - Sabrina S Burgener
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, QLD 4072, Australia
| | - Kate Schroder
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, QLD 4072, Australia
| | - Evan Ingley
- Cell Signalling Group, Harry Perkins Institute of Medical Research and Centre for Medical Research, The University of Western Australia, Perth, WA 6009, Australia; Discipline of Medical, Molecular and Forensic Sciences, College of Science, Health, Engineering and Education, Murdoch University, Murdoch, WA 6150, Australia
| | - Matthew J Sweet
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, QLD 4072, Australia
| | - Jennifer L Stow
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, QLD 4072, Australia.
| | - Lin Luo
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, QLD 4072, Australia.
| |
Collapse
|
|
20
|
Wang KL, Chen SN, Li L, Huo HJ, Nie P. Functional characterization of four TIR domain-containing adaptors, MyD88, TRIF, MAL, and SARM in mandarin fish Siniperca chuatsi. DEVELOPMENTAL AND COMPARATIVE IMMUNOLOGY 2021; 122:104110. [PMID: 33933533 DOI: 10.1016/j.dci.2021.104110] [Citation(s) in RCA: 5] [Impact Index Per Article: 1.7] [Reference Citation Analysis] [Abstract] [Key Words] [MESH Headings] [Track Full Text] [Subscribe] [Scholar Register] [Received: 02/08/2021] [Revised: 04/24/2021] [Accepted: 04/25/2021] [Indexed: 06/12/2023]
Abstract
Toll/interleukin-1 receptor (TIR) domain-containing adaptors, serve as pivotal signal transduction molecules in Toll-like receptor (TLR) signalling pathway to mediate downstream signalling cascades. In this study, four TIR-domain containing adaptors, MyD88, TRIF, MAL and SARM, were identified in mandarin fish Siniperca chuatsi, and they all contain TIR domains, of which MyD88 and SARM had high sequence homology with their vertebrate homologues. The expression analysis at mRNA level indicated that these genes were ubiquitously distributed in different tissues, being high in immune- and mucosa-related tissues such as head-kidney and intestine. The transcripts of these adaptor genes were up-regulated by poly(I:C) and LPS stimulation in isolated head-kidney lymphocytes (HKLs) of mandarin fish. Fluorescence microscopy revealed that all these molecules were localized in cytoplasm, and further investigations showed that the over-expression of MyD88, TRIF and MAL activated the NF-κB, ISRE or type Ι IFN promoters and inhibited SVCV replication, whereas their antiviral effects were significantly impaired when co-transfected with SARM. It was also confirmed by co-immunoprecipitation (Co-IP) that SARM interacts separately with MyD88, TRIF and MAL, and MAL interacts with MyD88. However, the regulatory mechanisms of these adaptors involved in signalling pathways of different TLRs should be of interest for further research.
Collapse
Affiliation(s)
- Kai Lun Wang
- State Key Laboratory of Freshwater Ecology and Biotechnology, And Key Laboratory of Aquaculture Disease Control, Institute of Hydrobiology, Chinese Academy of Sciences, Wuhan, Hubei Province, 430072, China; University of Chinese Academy of Sciences, Beijing, 100049, China; The Innovation Academy of Seed Design, Chinese Academy of Sciences, Wuhan, China
| | - Shan Nan Chen
- State Key Laboratory of Freshwater Ecology and Biotechnology, And Key Laboratory of Aquaculture Disease Control, Institute of Hydrobiology, Chinese Academy of Sciences, Wuhan, Hubei Province, 430072, China; The Innovation Academy of Seed Design, Chinese Academy of Sciences, Wuhan, China
| | - Li Li
- State Key Laboratory of Freshwater Ecology and Biotechnology, And Key Laboratory of Aquaculture Disease Control, Institute of Hydrobiology, Chinese Academy of Sciences, Wuhan, Hubei Province, 430072, China; The Innovation Academy of Seed Design, Chinese Academy of Sciences, Wuhan, China
| | - Hui Jun Huo
- Laboratory for Marine Biology and Biotechnology, Pilot National Laboratory for Marine Science and Technology (Qingdao), Qingdao, Shandong Province, 266237, China; School of Marine Science and Engineering, Qingdao Agricultural University, Qingdao, Shandong Province, 266109, China
| | - Pin Nie
- State Key Laboratory of Freshwater Ecology and Biotechnology, And Key Laboratory of Aquaculture Disease Control, Institute of Hydrobiology, Chinese Academy of Sciences, Wuhan, Hubei Province, 430072, China; The Innovation Academy of Seed Design, Chinese Academy of Sciences, Wuhan, China; Laboratory for Marine Biology and Biotechnology, Pilot National Laboratory for Marine Science and Technology (Qingdao), Qingdao, Shandong Province, 266237, China; School of Marine Science and Engineering, Qingdao Agricultural University, Qingdao, Shandong Province, 266109, China.
| |
Collapse
|
|
21
|
Nie X, Qian L, Sun R, Huang B, Dong X, Xiao Q, Zhang Q, Lu T, Yue L, Chen S, Li X, Sun Y, Li L, Xu L, Li Y, Yang M, Xue Z, Liang S, Ding X, Yuan C, Peng L, Liu W, Yi X, Lyu M, Xiao G, Xu X, Ge W, He J, Fan J, Wu J, Luo M, Chang X, Pan H, Cai X, Zhou J, Yu J, Gao H, Xie M, Wang S, Ruan G, Chen H, Su H, Mei H, Luo D, Zhao D, Xu F, Li Y, Zhu Y, Xia J, Hu Y, Guo T. Multi-organ proteomic landscape of COVID-19 autopsies. Cell 2021; 184:775-791.e14. [PMID: 33503446 PMCID: PMC7794601 DOI: 10.1016/j.cell.2021.01.004] [Citation(s) in RCA: 250] [Impact Index Per Article: 83.3] [Reference Citation Analysis] [Abstract] [Key Words] [Download PDF] [Figures] [Journal Information] [Subscribe] [Scholar Register] [Received: 08/16/2020] [Revised: 10/22/2020] [Accepted: 01/05/2021] [Indexed: 02/06/2023]
Abstract
The molecular pathology of multi-organ injuries in COVID-19 patients remains unclear, preventing effective therapeutics development. Here, we report a proteomic analysis of 144 autopsy samples from seven organs in 19 COVID-19 patients. We quantified 11,394 proteins in these samples, in which 5,336 were perturbed in the COVID-19 patients compared to controls. Our data showed that cathepsin L1, rather than ACE2, was significantly upregulated in the lung from the COVID-19 patients. Systemic hyperinflammation and dysregulation of glucose and fatty acid metabolism were detected in multiple organs. We also observed dysregulation of key factors involved in hypoxia, angiogenesis, blood coagulation, and fibrosis in multiple organs from the COVID-19 patients. Evidence for testicular injuries includes reduced Leydig cells, suppressed cholesterol biosynthesis, and sperm mobility. In summary, this study depicts a multi-organ proteomic landscape of COVID-19 autopsies that furthers our understanding of the biological basis of COVID-19 pathology.
Collapse
Affiliation(s)
- Xiu Nie
- Department of Pathology, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China
| | - Liujia Qian
- Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou 310024, China; Center for Infectious Disease Research, Westlake Laboratory of Life Sciences and Biomedicine, Hangzhou 310024, China; Institute of Basic Medical Sciences, Westlake Institute for Advanced Study, Hangzhou 310024, China
| | - Rui Sun
- Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou 310024, China; Center for Infectious Disease Research, Westlake Laboratory of Life Sciences and Biomedicine, Hangzhou 310024, China; Institute of Basic Medical Sciences, Westlake Institute for Advanced Study, Hangzhou 310024, China
| | - Bo Huang
- Department of Pathology, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China
| | - Xiaochuan Dong
- Department of Pathology, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China
| | - Qi Xiao
- Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou 310024, China; Center for Infectious Disease Research, Westlake Laboratory of Life Sciences and Biomedicine, Hangzhou 310024, China; Institute of Basic Medical Sciences, Westlake Institute for Advanced Study, Hangzhou 310024, China
| | - Qiushi Zhang
- Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou 310024, China; Center for Infectious Disease Research, Westlake Laboratory of Life Sciences and Biomedicine, Hangzhou 310024, China; Institute of Basic Medical Sciences, Westlake Institute for Advanced Study, Hangzhou 310024, China; Westlake Omics (Hangzhou) Biotechnology Co., Ltd., Hangzhou 310024, China
| | - Tian Lu
- Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou 310024, China; Center for Infectious Disease Research, Westlake Laboratory of Life Sciences and Biomedicine, Hangzhou 310024, China; Institute of Basic Medical Sciences, Westlake Institute for Advanced Study, Hangzhou 310024, China
| | - Liang Yue
- Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou 310024, China; Center for Infectious Disease Research, Westlake Laboratory of Life Sciences and Biomedicine, Hangzhou 310024, China; Institute of Basic Medical Sciences, Westlake Institute for Advanced Study, Hangzhou 310024, China
| | - Shuo Chen
- Department of Pathology, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China
| | - Xiang Li
- Department of Pathology, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China
| | - Yaoting Sun
- Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou 310024, China; Center for Infectious Disease Research, Westlake Laboratory of Life Sciences and Biomedicine, Hangzhou 310024, China; Institute of Basic Medical Sciences, Westlake Institute for Advanced Study, Hangzhou 310024, China
| | - Lu Li
- Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou 310024, China; Center for Infectious Disease Research, Westlake Laboratory of Life Sciences and Biomedicine, Hangzhou 310024, China; Institute of Basic Medical Sciences, Westlake Institute for Advanced Study, Hangzhou 310024, China
| | - Luang Xu
- Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou 310024, China; Center for Infectious Disease Research, Westlake Laboratory of Life Sciences and Biomedicine, Hangzhou 310024, China; Institute of Basic Medical Sciences, Westlake Institute for Advanced Study, Hangzhou 310024, China
| | - Yan Li
- Department of Pathology, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China
| | - Ming Yang
- Department of Pathology, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China
| | - Zhangzhi Xue
- Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou 310024, China; Center for Infectious Disease Research, Westlake Laboratory of Life Sciences and Biomedicine, Hangzhou 310024, China; Institute of Basic Medical Sciences, Westlake Institute for Advanced Study, Hangzhou 310024, China
| | - Shuang Liang
- Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou 310024, China; Center for Infectious Disease Research, Westlake Laboratory of Life Sciences and Biomedicine, Hangzhou 310024, China; Institute of Basic Medical Sciences, Westlake Institute for Advanced Study, Hangzhou 310024, China
| | - Xuan Ding
- Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou 310024, China; Center for Infectious Disease Research, Westlake Laboratory of Life Sciences and Biomedicine, Hangzhou 310024, China; Institute of Basic Medical Sciences, Westlake Institute for Advanced Study, Hangzhou 310024, China
| | - Chunhui Yuan
- Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou 310024, China; Center for Infectious Disease Research, Westlake Laboratory of Life Sciences and Biomedicine, Hangzhou 310024, China; Institute of Basic Medical Sciences, Westlake Institute for Advanced Study, Hangzhou 310024, China
| | - Li Peng
- Department of Pathology, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China
| | - Wei Liu
- Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou 310024, China; Center for Infectious Disease Research, Westlake Laboratory of Life Sciences and Biomedicine, Hangzhou 310024, China; Institute of Basic Medical Sciences, Westlake Institute for Advanced Study, Hangzhou 310024, China
| | - Xiao Yi
- Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou 310024, China; Center for Infectious Disease Research, Westlake Laboratory of Life Sciences and Biomedicine, Hangzhou 310024, China; Institute of Basic Medical Sciences, Westlake Institute for Advanced Study, Hangzhou 310024, China
| | - Mengge Lyu
- Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou 310024, China; Center for Infectious Disease Research, Westlake Laboratory of Life Sciences and Biomedicine, Hangzhou 310024, China; Institute of Basic Medical Sciences, Westlake Institute for Advanced Study, Hangzhou 310024, China
| | - Guixiang Xiao
- Department of Pathology, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China
| | - Xia Xu
- Department of Pathology, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China
| | - Weigang Ge
- Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou 310024, China; Center for Infectious Disease Research, Westlake Laboratory of Life Sciences and Biomedicine, Hangzhou 310024, China; Institute of Basic Medical Sciences, Westlake Institute for Advanced Study, Hangzhou 310024, China; Westlake Omics (Hangzhou) Biotechnology Co., Ltd., Hangzhou 310024, China
| | - Jiale He
- Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou 310024, China; Center for Infectious Disease Research, Westlake Laboratory of Life Sciences and Biomedicine, Hangzhou 310024, China; Institute of Basic Medical Sciences, Westlake Institute for Advanced Study, Hangzhou 310024, China
| | - Jun Fan
- Department of Pathology, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China
| | - Junhua Wu
- Department of Pathology, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China
| | - Meng Luo
- Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou 310024, China; Center for Infectious Disease Research, Westlake Laboratory of Life Sciences and Biomedicine, Hangzhou 310024, China; Institute of Basic Medical Sciences, Westlake Institute for Advanced Study, Hangzhou 310024, China; Department of Anatomy, College of Basic Medical Sciences, Dalian Medical University, Dalian 116044, China
| | - Xiaona Chang
- Department of Pathology, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China
| | - Huaxiong Pan
- Department of Pathology, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China
| | - Xue Cai
- Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou 310024, China; Center for Infectious Disease Research, Westlake Laboratory of Life Sciences and Biomedicine, Hangzhou 310024, China; Institute of Basic Medical Sciences, Westlake Institute for Advanced Study, Hangzhou 310024, China
| | - Junjie Zhou
- Department of Pathology, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China
| | - Jing Yu
- Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou 310024, China; Center for Infectious Disease Research, Westlake Laboratory of Life Sciences and Biomedicine, Hangzhou 310024, China; Institute of Basic Medical Sciences, Westlake Institute for Advanced Study, Hangzhou 310024, China
| | - Huanhuan Gao
- Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou 310024, China; Center for Infectious Disease Research, Westlake Laboratory of Life Sciences and Biomedicine, Hangzhou 310024, China; Institute of Basic Medical Sciences, Westlake Institute for Advanced Study, Hangzhou 310024, China
| | - Mingxing Xie
- Department of Ultrasound, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China
| | - Sihua Wang
- Department of Thoracic Surgery, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China
| | - Guan Ruan
- Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou 310024, China; Center for Infectious Disease Research, Westlake Laboratory of Life Sciences and Biomedicine, Hangzhou 310024, China; Institute of Basic Medical Sciences, Westlake Institute for Advanced Study, Hangzhou 310024, China
| | - Hao Chen
- Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou 310024, China; Center for Infectious Disease Research, Westlake Laboratory of Life Sciences and Biomedicine, Hangzhou 310024, China; Institute of Basic Medical Sciences, Westlake Institute for Advanced Study, Hangzhou 310024, China; Westlake Omics (Hangzhou) Biotechnology Co., Ltd., Hangzhou 310024, China
| | - Hua Su
- Department of Nephrology, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China
| | - Heng Mei
- Institute of Hematology, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China
| | - Danju Luo
- Department of Pathology, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China
| | - Dashi Zhao
- Department of Pathology, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China
| | - Fei Xu
- Department of Anatomy, College of Basic Medical Sciences, Dalian Medical University, Dalian 116044, China
| | - Yan Li
- Department of Anatomy and Physiology, College of Basic Medical Sciences, Shanghai Jiao Tong University, Shanghai 200025, China
| | - Yi Zhu
- Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou 310024, China; Center for Infectious Disease Research, Westlake Laboratory of Life Sciences and Biomedicine, Hangzhou 310024, China; Institute of Basic Medical Sciences, Westlake Institute for Advanced Study, Hangzhou 310024, China.
| | - Jiahong Xia
- Department of Cardiovascular Surgery, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China.
| | - Yu Hu
- Institute of Hematology, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China.
| | - Tiannan Guo
- Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou 310024, China; Center for Infectious Disease Research, Westlake Laboratory of Life Sciences and Biomedicine, Hangzhou 310024, China; Institute of Basic Medical Sciences, Westlake Institute for Advanced Study, Hangzhou 310024, China.
| |
Collapse
|
|
22
|
Kapetanovic R, Afroz SF, Ramnath D, Lawrence GM, Okada T, Curson JE, de Bruin J, Fairlie DP, Schroder K, St John JC, Blumenthal A, Sweet MJ. Lipopolysaccharide promotes Drp1-dependent mitochondrial fission and associated inflammatory responses in macrophages. Immunol Cell Biol 2020; 98:528-539. [PMID: 32686869 PMCID: PMC7497224 DOI: 10.1111/imcb.12363] [Citation(s) in RCA: 42] [Impact Index Per Article: 10.5] [Reference Citation Analysis] [Abstract] [Key Words] [Track Full Text] [Download PDF] [Figures] [Journal Information] [Subscribe] [Scholar Register] [Received: 04/08/2020] [Revised: 05/21/2020] [Accepted: 05/26/2020] [Indexed: 12/11/2022]
Abstract
Mitochondria have a multitude of functions, including energy generation and cell signaling. Recent evidence suggests that mitochondrial dynamics (i.e. the balance between mitochondrial fission and fusion) also regulate immune functions. Here, we reveal that lipopolysaccharide (LPS) stimulation increases mitochondrial numbers in mouse bone marrow‐derived macrophages (BMMs) and human monocyte‐derived macrophages. In BMMs, this response requires Toll‐like receptor 4 (Tlr4) and the TLR adaptor protein myeloid differentiation primary response 88 (MyD88) but is independent of mitochondrial biogenesis. Consistent with this phenomenon being a consequence of mitochondrial fission, the dynamin‐related protein 1 (Drp1) GTPase that promotes mitochondrial fission is enriched on mitochondria in LPS‐activated macrophages and is required for the LPS‐mediated increase in mitochondrial numbers in both BMMs and mouse embryonic fibroblasts. Pharmacological agents that skew toward mitochondrial fusion also abrogated this response. LPS triggered acute Drp1 phosphorylation at serine 635 (S635), followed by sustained Drp1 dephosphorylation at serine 656 (S656), in BMMs. LPS‐induced S656 dephosphorylation was abrogated in MyD88‐deficient BMMs, suggesting that this post‐translational modification is particularly important for Tlr4‐inducible fission. Pharmacological or genetic targeting of Tlr4‐inducible fission had selective effects on inflammatory mediator production, with LPS‐inducible mitochondrial fission promoting the expression and/or secretion of a subset of inflammatory mediators in BMMs and mouse embryonic fibroblasts. Thus, triggering of Tlr4 results in MyD88‐dependent activation of Drp1, leading to inducible mitochondrial fission and subsequent inflammatory responses in macrophages.
Collapse
Affiliation(s)
- Ronan Kapetanovic
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, QLD, 4072, Australia.,Australian Infectious Diseases Research Centre, The University of Queensland, Brisbane, QLD, 4072, Australia
| | - Syeda Farhana Afroz
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, QLD, 4072, Australia.,Australian Infectious Diseases Research Centre, The University of Queensland, Brisbane, QLD, 4072, Australia
| | - Divya Ramnath
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, QLD, 4072, Australia.,Australian Infectious Diseases Research Centre, The University of Queensland, Brisbane, QLD, 4072, Australia
| | - Grace Mep Lawrence
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, QLD, 4072, Australia.,Australian Infectious Diseases Research Centre, The University of Queensland, Brisbane, QLD, 4072, Australia
| | - Takashi Okada
- The Mitochondrial Genetics Group, Robinson Research Institute, School of Medicine, Adelaide Health and Medical Sciences Building, The University of Adelaide, Adelaide, SA, 5005, Australia
| | - James Eb Curson
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, QLD, 4072, Australia.,Australian Infectious Diseases Research Centre, The University of Queensland, Brisbane, QLD, 4072, Australia
| | - Jost de Bruin
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, QLD, 4072, Australia.,Australian Infectious Diseases Research Centre, The University of Queensland, Brisbane, QLD, 4072, Australia
| | - David P Fairlie
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, QLD, 4072, Australia.,Australian Infectious Diseases Research Centre, The University of Queensland, Brisbane, QLD, 4072, Australia
| | - Kate Schroder
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, QLD, 4072, Australia.,Australian Infectious Diseases Research Centre, The University of Queensland, Brisbane, QLD, 4072, Australia
| | - Justin C St John
- The Mitochondrial Genetics Group, Robinson Research Institute, School of Medicine, Adelaide Health and Medical Sciences Building, The University of Adelaide, Adelaide, SA, 5005, Australia
| | - Antje Blumenthal
- The University of Queensland Diamantina Institute, The University of Queensland, Brisbane, QLD, 4102, Australia
| | - Matthew J Sweet
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, QLD, 4072, Australia.,Australian Infectious Diseases Research Centre, The University of Queensland, Brisbane, QLD, 4072, Australia
| |
Collapse
|
|
23
|
Stocks CJ, von Pein JB, Curson JEB, Rae J, Phan MD, Foo D, Bokil NJ, Kambe T, Peters KM, Parton RG, Schembri MA, Kapetanovic R, Sweet MJ. Frontline Science: LPS-inducible SLC30A1 drives human macrophage-mediated zinc toxicity against intracellular Escherichia coli. J Leukoc Biol 2020; 109:287-297. [PMID: 32441444 PMCID: PMC7891337 DOI: 10.1002/jlb.2hi0420-160r] [Citation(s) in RCA: 8] [Impact Index Per Article: 2.0] [Reference Citation Analysis] [Abstract] [Key Words] [Track Full Text] [Download PDF] [Figures] [Journal Information] [Subscribe] [Scholar Register] [Received: 03/02/2020] [Revised: 04/22/2020] [Accepted: 04/28/2020] [Indexed: 12/12/2022] Open
Abstract
TLR-inducible zinc toxicity is an antimicrobial mechanism utilized by macrophages, however knowledge of molecular mechanisms mediating this response is limited. Here, we show that E. coli exposed to zinc stress within primary human macrophages reside in membrane-bound vesicular compartments. Since SLC30A zinc exporters can deliver zinc into the lumen of vesicles, we examined LPS-regulated mRNA expression of Slc30a/SLC30A family members in primary mouse and human macrophages. A number of these transporters were dynamically regulated in both cell populations. In human monocyte-derived macrophages, LPS strongly up-regulated SLC30A1 mRNA and protein expression. In contrast, SLC30A1 was not LPS-inducible in macrophage-like PMA-differentiated THP-1 cells. We therefore ectopically expressed SLC30A1 in these cells, finding that this was sufficient to promote zinc-containing vesicle formation. The response was similar to that observed following LPS stimulation. Ectopically expressed SLC30A1 localized to both the plasma membrane and intracellular zinc-containing vesicles within LPS-stimulated THP-1 cells. Inducible overexpression of SLC30A1 in THP-1 cells infected with the Escherichia coli K-12 strain MG1655 augmented the zinc stress response of intracellular bacteria and promoted clearance. Furthermore, in THP-1 cells infected with an MG1655 zinc stress reporter strain, all bacteria contained within SLC30A1-positive compartments were subjected to zinc stress. Thus, SLC30A1 marks zinc-containing compartments associated with TLR-inducible zinc toxicity in human macrophages, and its ectopic over-expression is sufficient to initiate this antimicrobial pathway in these cells. Finally, SLC30A1 silencing did not compromise E. coli clearance by primary human macrophages, suggesting that other zinc exporters may also contribute to the zinc toxicity response.
Collapse
Affiliation(s)
- Claudia J Stocks
- Institute for Molecular Bioscience (IMB), The University of Queensland, St. Lucia, Queensland, Australia.,IMB Centre for Inflammation and Disease Research, The University of Queensland, St. Lucia, Queensland, Australia.,Australian Infectious Diseases Research Centre, The University of Queensland, St. Lucia, Queensland, Australia
| | - Jessica B von Pein
- Institute for Molecular Bioscience (IMB), The University of Queensland, St. Lucia, Queensland, Australia.,IMB Centre for Inflammation and Disease Research, The University of Queensland, St. Lucia, Queensland, Australia.,Australian Infectious Diseases Research Centre, The University of Queensland, St. Lucia, Queensland, Australia
| | - James E B Curson
- Institute for Molecular Bioscience (IMB), The University of Queensland, St. Lucia, Queensland, Australia.,IMB Centre for Inflammation and Disease Research, The University of Queensland, St. Lucia, Queensland, Australia.,Australian Infectious Diseases Research Centre, The University of Queensland, St. Lucia, Queensland, Australia
| | - James Rae
- Institute for Molecular Bioscience (IMB), The University of Queensland, St. Lucia, Queensland, Australia
| | - Minh-Duy Phan
- Australian Infectious Diseases Research Centre, The University of Queensland, St. Lucia, Queensland, Australia.,School of Chemistry and Molecular Biosciences, The University of Queensland, St. Lucia, Queensland, Australia
| | - Darren Foo
- Institute for Molecular Bioscience (IMB), The University of Queensland, St. Lucia, Queensland, Australia.,IMB Centre for Inflammation and Disease Research, The University of Queensland, St. Lucia, Queensland, Australia.,Australian Infectious Diseases Research Centre, The University of Queensland, St. Lucia, Queensland, Australia
| | - Nilesh J Bokil
- Institute for Molecular Bioscience (IMB), The University of Queensland, St. Lucia, Queensland, Australia.,IMB Centre for Inflammation and Disease Research, The University of Queensland, St. Lucia, Queensland, Australia.,Australian Infectious Diseases Research Centre, The University of Queensland, St. Lucia, Queensland, Australia
| | - Taiho Kambe
- Division of Integrated Life Science, Graduate School of Biostudies, Kyoto University, Kyoto, Japan
| | - Kate M Peters
- Australian Infectious Diseases Research Centre, The University of Queensland, St. Lucia, Queensland, Australia.,School of Chemistry and Molecular Biosciences, The University of Queensland, St. Lucia, Queensland, Australia
| | - Robert G Parton
- Institute for Molecular Bioscience (IMB), The University of Queensland, St. Lucia, Queensland, Australia.,Centre for Microscopy and Microanalysis, The University of Queensland, St. Lucia, Queensland, Australia
| | - Mark A Schembri
- Australian Infectious Diseases Research Centre, The University of Queensland, St. Lucia, Queensland, Australia.,School of Chemistry and Molecular Biosciences, The University of Queensland, St. Lucia, Queensland, Australia
| | - Ronan Kapetanovic
- Institute for Molecular Bioscience (IMB), The University of Queensland, St. Lucia, Queensland, Australia.,IMB Centre for Inflammation and Disease Research, The University of Queensland, St. Lucia, Queensland, Australia.,Australian Infectious Diseases Research Centre, The University of Queensland, St. Lucia, Queensland, Australia
| | - Matthew J Sweet
- Institute for Molecular Bioscience (IMB), The University of Queensland, St. Lucia, Queensland, Australia.,IMB Centre for Inflammation and Disease Research, The University of Queensland, St. Lucia, Queensland, Australia.,Australian Infectious Diseases Research Centre, The University of Queensland, St. Lucia, Queensland, Australia
| |
Collapse
|
|
24
|
He Y, Ji P, Li Y, Wang R, Ma H, Yuan H. Genetic Variants Were Associated With the Prognosis of Head and Neck Squamous Carcinoma. Front Oncol 2020; 10:372. [PMID: 32266149 PMCID: PMC7099049 DOI: 10.3389/fonc.2020.00372] [Citation(s) in RCA: 6] [Impact Index Per Article: 1.5] [Reference Citation Analysis] [Abstract] [Key Words] [Track Full Text] [Download PDF] [Figures] [Journal Information] [Subscribe] [Scholar Register] [Received: 09/27/2019] [Accepted: 03/03/2020] [Indexed: 12/12/2022] Open
Abstract
Background: As the sixth most common cancer of worldwide, head and neck cancers (HNC) are springing from oral cavity, pharynx and larynx and there is no strong biomarker for prognosis. Rates of 5 years survival with HNC remain relatively low in decades with improvement of treatments. Evidence that single nucleotide polymorphisms (SNPs) play a part in cancer prognosis is growing. Methods: We conducted an exome-wide association study among 261 patients with head and neck squamous cell carcinoma (HNSCC) and then validated in The Cancer Genome Atlas (TCGA) database for survival by using the Cox proportional hazards regression models and Kaplan–Meier analyses. Results: After combining the result of the two stages, 4 SNPs were significantly associated with HNSCC survival (rs16879870 at 6q14.3: adjusted HR = 2.02, 95%CI = 1.50–2.73, P = 3.88 × 10−6; rs2641256 at 17p13.2: adjusted HR = 0.67, 95%CI = 0.56–0.80, P = 7.51 × 10−6; rs2761591 at 11p13: adjusted HR = 2.07, 95%CI = 1.50–2.87, P = 1.16 × 10−5; and rs854936 at 22q11.21: adjusted HR = 1.92, 95%CI = 1.43–2.57, P = 1.27 × 10−5). Besides, we constructed a receiver operating characteristic (ROC) model to estimate predictive effect of the novel SNPs combined with clinical stage in HNSCC prognosis (AUC = 0.715). We also found the genotype of rs16879870 and rs854936 was significantly associated with the expression of gene GJB7 (P = 0.013) and RTN4R (P = 0.047) in cancer tissues of TCGA, respectively. Conclusion: Our findings suggested that the SNPs (rs16879870, rs2641256, rs2761591, rs854936) might play a crucial role in prognosis of HNSCC.
Collapse
Affiliation(s)
- Yingzheng He
- Jiangsu Key Laboratory of Oral Diseases, Nanjing Medical University, Nanjing, China.,Department of Oral and Maxillofacial Surgery, Affiliated Hospital of Stomatology, Nanjing Medical University, Nanjing, China
| | - Pei Ji
- Jiangsu Key Laboratory of Cancer Biomarkers, Prevention and Treatment, Cancer Center, Department of Epidemiology and Biostatistics, School of Public Health, Nanjing Medical University, Nanjing, China
| | - Yuancheng Li
- Jiangsu Key Laboratory of Cancer Biomarkers, Prevention and Treatment, Cancer Center, Department of Epidemiology and Biostatistics, School of Public Health, Nanjing Medical University, Nanjing, China
| | - Ruixia Wang
- Jiangsu Key Laboratory of Oral Diseases, Nanjing Medical University, Nanjing, China
| | - Hongxia Ma
- Jiangsu Key Laboratory of Cancer Biomarkers, Prevention and Treatment, Cancer Center, Department of Epidemiology and Biostatistics, School of Public Health, Nanjing Medical University, Nanjing, China
| | - Hua Yuan
- Jiangsu Key Laboratory of Oral Diseases, Nanjing Medical University, Nanjing, China.,Department of Oral and Maxillofacial Surgery, Affiliated Hospital of Stomatology, Nanjing Medical University, Nanjing, China
| |
Collapse
|
|
25
|
Alzheimer’s Disease Genetics: Review of Novel Loci Associated with Disease. CURRENT GENETIC MEDICINE REPORTS 2020. [DOI: 10.1007/s40142-020-00182-y] [Citation(s) in RCA: 10] [Impact Index Per Article: 2.5] [Reference Citation Analysis] [Track Full Text] [Journal Information] [Subscribe] [Scholar Register] [Indexed: 12/16/2022]
|
|
26
|
Ullah MA, Vicente CT, Collinson N, Curren B, Sikder MAA, Sebina I, Simpson J, Varelias A, Lindquist JA, Ferreira MAR, Phipps S. PAG1 limits allergen-induced type 2 inflammation in the murine lung. Allergy 2020; 75:336-345. [PMID: 31321783 DOI: 10.1111/all.13991] [Citation(s) in RCA: 9] [Impact Index Per Article: 2.3] [Reference Citation Analysis] [Abstract] [Key Words] [Track Full Text] [Journal Information] [Subscribe] [Scholar Register] [Received: 03/31/2019] [Revised: 05/30/2019] [Accepted: 06/24/2019] [Indexed: 12/24/2022]
Abstract
BACKGROUND Phosphoprotein associated with glycosphingolipid-enriched microdomains 1 (PAG1) is a transmembrane adaptor protein that affects immune receptor signaling in T and B cells. Evidence from genome-wide association studies of asthma suggests that genetic variants that regulate the expression of PAG1 are associated with asthma risk. However, it is not known whether PAG1 expression is causally related to asthma pathophysiology. Here, we investigated the role of PAG1 in a preclinical mouse model of house dust mite (HDM)-induced allergic sensitization and allergic airway inflammation. METHODS Pag1-deficient (Pag1-/- ) and wild-type (WT) mice were sensitized or sensitized/challenged to HDM, and hallmark features of allergic inflammation were assessed. The contribution of T cells was assessed through depletion (anti-CD4 antibody) and adoptive transfer studies. RESULTS Type 2 inflammation (eosinophilia, eotaxin-2 expression, IL-4/IL-5/IL-13 production, mucus production) in the airways and lungs was significantly increased in HDM sensitized/challenged Pag1-/- mice compared to WT mice. The predisposition to allergic sensitization was associated with increased airway epithelial high-mobility group box 1 (HMGB1) translocation and release, increased type 2 innate lymphoid cells (ILC2s) and monocyte-derived dendritic cell numbers in the mediastinal lymph nodes, and increased T-helper type 2 (TH 2)-cell differentiation. CD4+ T-cell depletion studies or the adoptive transfer of WT OVA-specific CD4+ T cells to WT or Pag1-/- recipients demonstrated that the heightened propensity for TH 2-cell differentiation was both T cell intrinsic and extrinsic. CONCLUSION PAG1 deficiency increased airway epithelial activation, ILC2 expansion, and TH 2 differentiation. As a consequence, PAG1 deficiency predisposed toward allergic sensitization and increased the severity of experimental asthma.
Collapse
Affiliation(s)
- Md Ashik Ullah
- QIMR Berghofer Medical Research Institute Brisbane Qld Australia
- Faculty of Medicine University of Queensland Brisbane Qld Australia
| | - Cristina T. Vicente
- QIMR Berghofer Medical Research Institute Brisbane Qld Australia
- Faculty of Medicine University of Queensland Brisbane Qld Australia
| | | | - Bodie Curren
- QIMR Berghofer Medical Research Institute Brisbane Qld Australia
- Faculty of Medicine University of Queensland Brisbane Qld Australia
| | - Md Al Amin Sikder
- QIMR Berghofer Medical Research Institute Brisbane Qld Australia
- Faculty of Medicine University of Queensland Brisbane Qld Australia
| | - Ismail Sebina
- QIMR Berghofer Medical Research Institute Brisbane Qld Australia
| | - Jennifer Simpson
- QIMR Berghofer Medical Research Institute Brisbane Qld Australia
- Faculty of Medicine University of Queensland Brisbane Qld Australia
| | - Antiopi Varelias
- QIMR Berghofer Medical Research Institute Brisbane Qld Australia
- Faculty of Medicine University of Queensland Brisbane Qld Australia
| | - Jonathan A. Lindquist
- Clinic for Nephrology and Hypertension, Diabetology and Endocrinology Otto‐von‐Guericke University Magdeburg Germany
| | | | - Simon Phipps
- QIMR Berghofer Medical Research Institute Brisbane Qld Australia
- Faculty of Medicine University of Queensland Brisbane Qld Australia
- Australian Infectious Diseases Research Centre University of Queensland Brisbane Qld Australia
| |
Collapse
|
|
27
|
Wall AA, Condon ND, Luo L, Stow JL. Rab8a localisation and activation by Toll-like receptors on macrophage macropinosomes. Philos Trans R Soc Lond B Biol Sci 2020; 374:20180151. [PMID: 30966999 DOI: 10.1098/rstb.2018.0151] [Citation(s) in RCA: 18] [Impact Index Per Article: 4.5] [Reference Citation Analysis] [Abstract] [Key Words] [Track Full Text] [Journal Information] [Subscribe] [Scholar Register] [Indexed: 12/11/2022] Open
Abstract
Macropinocytosis is a prevalent and essential pathway in macrophages where it contributes to anti-microbial responses and innate immune cell functions. Cell surface ruffles give rise to phagosomes and to macropinosomes as multi-functional compartments that contribute to environmental sampling, pathogen entry, plasma membrane turnover and receptor signalling. Rapid, high resolution, lattice light sheet imaging demonstrates the dynamic nature of macrophage ruffling. Pathogen-mediated activation of surface and endosomal Toll-like receptors (TLRs) in macrophages upregulates macropinocytosis. Here, using multiple forms of imaging and microscopy, we track membrane-associated, fluorescently-tagged Rab8a expressed in live macrophages, using a variety of cell markers to demonstrate Rab8a localization and its enrichment on early macropinosomes. Production of a novel biosensor and its use for quantitative FRET analysis in live cells, pinpoints macropinosomes as the site for TLR-induced activation of Rab8a. We have previously shown that TLR signalling, cytokine outputs and macrophage programming are regulated by the GTPase Rab8a with PI3 Kγ as its effector. Finally, we highlight another effector, the phosphatase OCRL, which is located on macropinosomes and interacts with Rab8a, suggesting that Rab8a may operate on multiple levels to modulate phosphoinositides in macropinosomes. These findings extend our understanding of macropinosomes as regulatory compartments for innate immune function in macrophages. This article is part of the Theo Murphy meeting issue 'Macropinocytosis'.
Collapse
Affiliation(s)
- Adam A Wall
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research, University of Queensland , Brisbane, Queensland 4072 , Australia
| | - Nicholas D Condon
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research, University of Queensland , Brisbane, Queensland 4072 , Australia
| | - Lin Luo
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research, University of Queensland , Brisbane, Queensland 4072 , Australia
| | - Jennifer L Stow
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research, University of Queensland , Brisbane, Queensland 4072 , Australia
| |
Collapse
|
|
28
|
Apolipoprotein C3 induces inflammation and organ damage by alternative inflammasome activation. Nat Immunol 2020; 21:30-41. [PMID: 31819254 DOI: 10.1038/s41590-019-0548-1] [Citation(s) in RCA: 154] [Impact Index Per Article: 38.5] [Reference Citation Analysis] [Abstract] [Track Full Text] [Journal Information] [Subscribe] [Scholar Register] [Received: 01/14/2019] [Accepted: 10/29/2019] [Indexed: 02/06/2023]
Abstract
NLRP3-inflammasome-driven inflammation is involved in the pathogenesis of a variety of diseases. Identification of endogenous inflammasome activators is essential for the development of new anti-inflammatory treatment strategies. Here, we identified that apolipoprotein C3 (ApoC3) activates the NLRP3 inflammasome in human monocytes by inducing an alternative NLRP3 inflammasome via caspase-8 and dimerization of Toll-like receptors 2 and 4. Alternative inflammasome activation in human monocytes is mediated by the Toll-like receptor adapter protein SCIMP. This triggers Lyn/Syk-dependent calcium entry and the production of reactive oxygen species, leading to activation of caspase-8. In humanized mouse models, ApoC3 activated human monocytes in vivo to impede endothelial regeneration and promote kidney injury in an NLRP3- and caspase-8-dependent manner. These data provide new insights into the regulation of the NLRP3 inflammasome and the pathophysiological role of triglyceride-rich lipoproteins containing ApoC3. Targeting ApoC3 might prevent organ damage and provide an anti-inflammatory treatment for vascular and kidney diseases.
Collapse
|
|
29
|
Luo L, Lucas RM, Liu L, Stow JL. Signalling, sorting and scaffolding adaptors for Toll-like receptors. J Cell Sci 2019; 133:133/5/jcs239194. [PMID: 31889021 DOI: 10.1242/jcs.239194] [Citation(s) in RCA: 52] [Impact Index Per Article: 10.4] [Reference Citation Analysis] [Abstract] [Key Words] [Track Full Text] [Journal Information] [Subscribe] [Scholar Register] [Indexed: 12/11/2022] Open
Abstract
Toll-like receptors (TLRs) are danger-sensing receptors that typically propagate self-limiting inflammatory responses, but can unleash uncontrolled inflammation in non-homeostatic or disease settings. Activation of TLRs by pathogen- and/or host-derived stimuli triggers a range of signalling and transcriptional pathways to programme inflammatory and anti-microbial responses, including the production of a suite of inflammatory cytokines and other mediators. Multiple sorting and signalling adaptors are recruited to receptor complexes on the plasma membrane or endosomes where they act as scaffolds for downstream signalling kinases and effectors at these sites. So far, seven proximal TLR adaptors have been identified: MyD88, MAL, TRIF (also known as TICAM1), TRAM (TICAM2), SARM (SARM1), BCAP (PIK3AP1) and SCIMP. Most adaptors tether directly to TLRs through homotypic Toll/interleukin-1 receptor domain (TIR)-TIR interactions, whereas SCIMP binds to TLRs through an atypical TIR-non-TIR interaction. In this Review, we highlight the key roles for these adaptors in TLR signalling, scaffolding and receptor sorting and discuss how the adaptors thereby direct the differential outcomes of TLR-mediated responses. We further summarise TLR adaptor regulation and function, and make note of human diseases that might be associated with mutations in these adaptors.
Collapse
Affiliation(s)
- Lin Luo
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, QLD 4072, Australia
| | - Richard M Lucas
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, QLD 4072, Australia
| | - Liping Liu
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, QLD 4072, Australia
| | - Jennifer L Stow
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research, The University of Queensland, Brisbane, QLD 4072, Australia
| |
Collapse
|
|
30
|
TLR Crosstalk Activates LRP1 to Recruit Rab8a and PI3Kγ for Suppression of Inflammatory Responses. Cell Rep 2019; 24:3033-3044. [PMID: 30208326 DOI: 10.1016/j.celrep.2018.08.028] [Citation(s) in RCA: 57] [Impact Index Per Article: 11.4] [Reference Citation Analysis] [Abstract] [Key Words] [Track Full Text] [Journal Information] [Subscribe] [Scholar Register] [Received: 01/04/2018] [Revised: 06/28/2018] [Accepted: 08/10/2018] [Indexed: 02/06/2023] Open
Abstract
The multi-ligand endocytic receptor, low-density lipoprotein-receptor-related protein 1 (LRP1), has anti-inflammatory roles in disease. Here, we reveal that pathogen-activated Toll-like receptors (TLRs) activate LRP1 in human and mouse primary macrophages, resulting in phosphorylation of LRP1 at Y4507. In turn, this allows LRP1 to activate and recruit the guanosine triphosphatase (GTPase), Rab8a, with p110γ/p101 as its phosphatidylinositol 3-kinase (PI3K) effector complex. PI3Kγ is a known regulator of TLR signaling and macrophage reprogramming. LRP1 coincides with Rab8a at signaling sites on macropinosomal membranes. In LRP1-deficient cells, TLR-induced Rab8 activation is abolished. CRISPR-mediated knockout of LRP1 in macrophages alters Akt/mTOR signaling and produces a pro-inflammatory bias in cytokine outputs, mimicking the Rab8a knockout and PI3Kγ-null phenotype. Thus, TLR-LRP1 crosstalk activates the Rab8a/PI3Kγ complex for reprogramming macrophages, revealing this as a key mechanism through which LRP1 helps to suppress inflammation.
Collapse
|
|
31
|
Ying Z, Li X, Dang H, Yin N, Gao C. Molecular immune mechanisms of HPV-infected HaCaT cells in vitro based on toll-like receptors signaling pathway. J Clin Lab Anal 2019; 34:e23101. [PMID: 31785031 PMCID: PMC7083446 DOI: 10.1002/jcla.23101] [Citation(s) in RCA: 2] [Impact Index Per Article: 0.4] [Reference Citation Analysis] [Abstract] [Key Words] [Track Full Text] [Download PDF] [Figures] [Journal Information] [Subscribe] [Scholar Register] [Received: 07/09/2019] [Revised: 09/09/2019] [Accepted: 09/11/2019] [Indexed: 12/19/2022] Open
Abstract
OBJECTIVE To explore the molecular immune mechanism of HPV-infected HaCaT cells in vitro based on TLRs signaling pathway by analyzing the effects of interfering TLRs on inflammatory and immune factors in the signaling pathway. METHODS FCM was used to analyze the proportion of Th1, Th2, Th17, and Treg cells in blood samples. HPV-infected HaCaT cells were divided into five groups: A, B, C, D, and E. Group A added TLR3 antagonist, group B added TLR9 antagonist, group C added equivalent saline, group D added IRF3 agonist, and group E added IRF3 inhibitor. Immunohistochemistry was used to analyze the expression of TLR3 and TLR9 in HaCaT cell model; ELISA was used to analyze the expression of inflammatory factors IL-2, TNF-a, and IFN-beta; WB was used to analyze the expression of TRAF3, IKK epsilon, and TBK1; RT-PCR was used to analyze the expression of IRF3 and IRF7 in each cell model. RESULTS The proportion of blood immune cells in patients with HPV infection was Th1, Th17, Th2, and Treg, with statistical significance (P < .05); the expression of TLR3 and TLR9 in HPV-infected cells was higher than that in negative control group, with statistical significance (P < .05); TLR3 was higher than TLR9, with no significant difference (P > .05); the expression of IL-2, TNF-alpha, IFN-beta in each group, TLR3, and TLR9 was higher than that in negative control group (P < .05). The expression of TRAF3, IKK epsilon, and TBK1 in the control group was higher than that in the TLR3 and TLR9 inhibitor groups, and the expression of IRF3 and IRF7 in the TLR9 inhibitor group was higher than that in the TLR3 inhibitor group (P < .05); the expression of IRF3 and IRF7 in the TLR3i and TLR9i inhibitor groups was lower than that in the TLR3 inhibitor group (P < .05). Compared with the control group, IRF3a group was higher than the control group, IRF3i group was lower than the control group, the difference was statistically significant (P < .05). CONCLUSION TLR3 and TLR9, the key factors of TLRs, are highly expressed in HaCaT cells infected with HPV. Through TLRs-IKK-e-IRFs-IFN signaling pathway, they can induce high expression of inflammatory factors, IKK-e, IRFs, and IFN, and improve immunity.
Collapse
Affiliation(s)
- Zuolin Ying
- Department of Dermatology, Shanghai General Hospital, Shanghai Jiao Tong University School of Medicine, Shanghai, China
| | - Xiaojie Li
- Department of Dermatology, Shanghai General Hospital, Shanghai Jiao Tong University School of Medicine, Shanghai, China
| | - Hong Dang
- Department of Dermatology, Shanghai General Hospital, Shanghai Jiao Tong University School of Medicine, Shanghai, China
| | - Na Yin
- Department of Dermatology, Shanghai General Hospital, Shanghai Jiao Tong University School of Medicine, Shanghai, China
| | - Chuang Gao
- Department of Dermatology, Shanghai General Hospital, Shanghai Jiao Tong University School of Medicine, Shanghai, China
| |
Collapse
|
|
32
|
Lauenstein JU, Udgata A, Bartram A, De Sutter D, Fisher DI, Halabi S, Eyckerman S, Gay NJ. Phosphorylation of the multifunctional signal transducer B-cell adaptor protein (BCAP) promotes recruitment of multiple SH2/SH3 proteins including GRB2. J Biol Chem 2019; 294:19852-19861. [PMID: 31527084 PMCID: PMC6937578 DOI: 10.1074/jbc.ra119.009931] [Citation(s) in RCA: 5] [Impact Index Per Article: 1.0] [Reference Citation Analysis] [Abstract] [Key Words] [Track Full Text] [Download PDF] [Figures] [Journal Information] [Subscribe] [Scholar Register] [Received: 06/28/2019] [Revised: 09/10/2019] [Indexed: 12/21/2022] Open
Abstract
B-cell adaptor protein (BCAP) is a multimodular, multifunctional signal transducer that regulates signal transduction pathways in leukocytes, including macrophages, B-cells, and T-cells. In particular, BCAP suppresses inflammatory signaling by Toll-like receptors (TLRs). However, how BCAP itself is regulated and what its interaction partners are is unclear. Here, using human immune cell lines, including THP-1 cells, we characterized the complex phosphorylation patterns of BCAP and used a novel protein complex trapping strategy, called virotrap, to identify its interaction partners. This analysis identified known interactions of BCAP with phosphoinositide 3-kinase (PI3K) p85 subunit and NCK adaptor protein (NCK), together with previously unknown interactions of BCAP with Src homology 2 (SH2) and SH3 domain-containing adaptor proteins, notably growth factor receptor-bound protein 2 (GRB2) and CRK-like proto-oncogene, adaptor protein (CRKL). We show that the SH3 domain of GRB2 can bind to BCAP independently of BCAP phosphorylation status, suggesting that the SH2 domains mediate interactions with activated receptor tyrosine kinase complexes including the CD19 subunit of the B-cell receptor. Our results also suggested that the PI3K p85 subunit binds to BCAP via SH3 domains forming an inactive complex that is then activated by sequential binding with the SH2 domains. Taken together, our results indicate that BCAP is a complex hub that processes signals from multiple pathways in diverse cell types of the immune system.
Collapse
Affiliation(s)
- Johannes U Lauenstein
- Department of Biochemistry, University of Cambridge, Cambridge CB2 1GA, United Kingdom
| | - Atul Udgata
- Department of Biochemistry, University of Cambridge, Cambridge CB2 1GA, United Kingdom
| | - Alex Bartram
- Department of Biochemistry, University of Cambridge, Cambridge CB2 1GA, United Kingdom
| | - Delphine De Sutter
- Department of Biomolecular Medicine, Ghent University, VIB Center for Medical Biotechnology, VIB, A. Baertsoenkaai 3, Ghent B-9000, Belgium
| | - David I Fisher
- Discovery Sciences, Discovery Biology, IMED Biotech Unit, AstraZeneca, Cambridge CB4 0WG, United Kingdom
| | - Samer Halabi
- Department of Biochemistry, University of Cambridge, Cambridge CB2 1GA, United Kingdom
| | - Sven Eyckerman
- Department of Biomolecular Medicine, Ghent University, VIB Center for Medical Biotechnology, VIB, A. Baertsoenkaai 3, Ghent B-9000, Belgium
| | - Nicholas J Gay
- Department of Biochemistry, University of Cambridge, Cambridge CB2 1GA, United Kingdom
| |
Collapse
|
|
33
|
The metabolic regulator Lamtor5 suppresses inflammatory signaling via regulating mTOR-mediated TLR4 degradation. Cell Mol Immunol 2019; 17:1063-1076. [PMID: 31467416 PMCID: PMC7608472 DOI: 10.1038/s41423-019-0281-6] [Citation(s) in RCA: 21] [Impact Index Per Article: 4.2] [Reference Citation Analysis] [Abstract] [Key Words] [Track Full Text] [Download PDF] [Figures] [Journal Information] [Subscribe] [Scholar Register] [Received: 08/11/2019] [Accepted: 08/13/2019] [Indexed: 01/10/2023] Open
Abstract
Comprehensive immune responses are essential for eliminating pathogens but must be tightly controlled to avoid sustained immune activation and potential tissue damage. The engagement of TLR4, a canonical pattern recognition receptor, has been proposed to trigger inflammatory responses with different magnitudes and durations depending on TLR4 cellular compartmentalization. In the present study, we identify an unexpected role of Lamtor5, a newly identified component of the amino acid-sensing machinery, in modulating TLR4 signaling and controlling inflammation. Specifically, Lamtor5 associated with TLR4 via their LZ/TIR domains and facilitated their colocalization at autolysosomes, preventing lysosomal tethering and the activation of mTORC1 upon LPS stimulation and thereby derepressing TFEB to promote autophagic degradation of TLR4. The loss of Lamtor5 was unable to trigger the TFEB-driven autolysosomal pathway and delay degradation of TLR4, leading to sustained inflammation and hence increased mortality among Lamtor5 haploinsufficient mice during endotoxic shock. Intriguingly, nutrient deprivation, particularly leucine deprivation, blunted inflammatory signaling and conferred protection to endotoxic mice. This effect, however, was largely abrogated upon Lamtor5 deletion. We thus propose a homeostatic function of Lamtor5 that couples pathogenic insults and nutrient availability to optimize the inflammatory response; this function may have implications for TLR4-associated inflammatory and metabolic disorders.
Collapse
|
|
34
|
Luo L, Curson JEB, Liu L, Wall AA, Tuladhar N, Lucas RM, Sweet MJ, Stow JL. SCIMP is a universal Toll‐like receptor adaptor in macrophages. J Leukoc Biol 2019; 107:251-262. [DOI: 10.1002/jlb.2ma0819-138rr] [Citation(s) in RCA: 9] [Impact Index Per Article: 1.8] [Reference Citation Analysis] [Track Full Text] [Journal Information] [Subscribe] [Scholar Register] [Received: 05/12/2019] [Revised: 08/19/2019] [Accepted: 08/20/2019] [Indexed: 02/06/2023] Open
Affiliation(s)
- Lin Luo
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research The University of Queensland Brisbane Queensland Australia
| | - James E. B. Curson
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research The University of Queensland Brisbane Queensland Australia
| | - Liping Liu
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research The University of Queensland Brisbane Queensland Australia
| | - Adam A. Wall
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research The University of Queensland Brisbane Queensland Australia
| | - Neeraj Tuladhar
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research The University of Queensland Brisbane Queensland Australia
| | - Richard M. Lucas
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research The University of Queensland Brisbane Queensland Australia
| | - Matthew J. Sweet
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research The University of Queensland Brisbane Queensland Australia
| | - Jennifer L. Stow
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research The University of Queensland Brisbane Queensland Australia
| |
Collapse
|
|
35
|
Moreno-Grau S, de Rojas I, Hernández I, Quintela I, Montrreal L, Alegret M, Hernández-Olasagarre B, Madrid L, González-Perez A, Maroñas O, Rosende-Roca M, Mauleón A, Vargas L, Lafuente A, Abdelnour C, Rodríguez-Gómez O, Gil S, Santos-Santos MÁ, Espinosa A, Ortega G, Sanabria Á, Pérez-Cordón A, Cañabate P, Moreno M, Preckler S, Ruiz S, Aguilera N, Pineda JA, Macías J, Alarcón-Martín E, Sotolongo-Grau O, Marquié M, Monté-Rubio G, Valero S, Benaque A, Clarimón J, Bullido MJ, García-Ribas G, Pástor P, Sánchez-Juan P, Álvarez V, Piñol-Ripoll G, García-Alberca JM, Royo JL, Franco E, Mir P, Calero M, Medina M, Rábano A, Ávila J, Antúnez C, Real LM, Orellana A, Carracedo Á, Sáez ME, Tárraga L, Boada M, Ruiz A. Genome-wide association analysis of dementia and its clinical endophenotypes reveal novel loci associated with Alzheimer's disease and three causality networks: The GR@ACE project. Alzheimers Dement 2019; 15:1333-1347. [PMID: 31473137 DOI: 10.1016/j.jalz.2019.06.4950] [Citation(s) in RCA: 91] [Impact Index Per Article: 18.2] [Reference Citation Analysis] [Abstract] [Key Words] [Track Full Text] [Journal Information] [Subscribe] [Scholar Register] [Received: 01/29/2019] [Revised: 06/19/2019] [Accepted: 06/19/2019] [Indexed: 01/10/2023]
Abstract
INTRODUCTION Large variability among Alzheimer's disease (AD) cases might impact genetic discoveries and complicate dissection of underlying biological pathways. METHODS Genome Research at Fundacio ACE (GR@ACE) is a genome-wide study of dementia and its clinical endophenotypes, defined based on AD's clinical certainty and vascular burden. We assessed the impact of known AD loci across endophenotypes to generate loci categories. We incorporated gene coexpression data and conducted pathway analysis per category. Finally, to evaluate the effect of heterogeneity in genetic studies, GR@ACE series were meta-analyzed with additional genome-wide association study data sets. RESULTS We classified known AD loci into three categories, which might reflect the disease clinical heterogeneity. Vascular processes were only detected as a causal mechanism in probable AD. The meta-analysis strategy revealed the ANKRD31-rs4704171 and NDUFAF6-rs10098778 and confirmed SCIMP-rs7225151 and CD33-rs3865444. DISCUSSION The regulation of vasculature is a prominent causal component of probable AD. GR@ACE meta-analysis revealed novel AD genetic signals, strongly driven by the presence of clinical heterogeneity in the AD series.
Collapse
Affiliation(s)
- Sonia Moreno-Grau
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain; CIBERNED, Network Center for Biomedical Research in Neurodegenerative Diseases, National Institute of Health Carlos III, Madrid, Spain
| | - Itziar de Rojas
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain
| | - Isabel Hernández
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain; CIBERNED, Network Center for Biomedical Research in Neurodegenerative Diseases, National Institute of Health Carlos III, Madrid, Spain
| | - Inés Quintela
- Grupo de Medicina Xenómica, Centro Nacional de Genotipado (CEGEN-PRB3-ISCIII). Universidade de Santiago de Compostela, Santiago de Compostela, Spain
| | - Laura Montrreal
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain
| | - Montserrat Alegret
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain; CIBERNED, Network Center for Biomedical Research in Neurodegenerative Diseases, National Institute of Health Carlos III, Madrid, Spain
| | - Begoña Hernández-Olasagarre
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain
| | - Laura Madrid
- CAEBI, Centro Andaluz de Estudios Bioinformáticos, Sevilla, Spain
| | | | - Olalla Maroñas
- Grupo de Medicina Xenómica, Centro Nacional de Genotipado (CEGEN-PRB3-ISCIII). Universidade de Santiago de Compostela, Santiago de Compostela, Spain
| | - Maitée Rosende-Roca
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain
| | - Ana Mauleón
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain
| | - Liliana Vargas
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain
| | - Asunción Lafuente
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain
| | - Carla Abdelnour
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain; CIBERNED, Network Center for Biomedical Research in Neurodegenerative Diseases, National Institute of Health Carlos III, Madrid, Spain
| | - Octavio Rodríguez-Gómez
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain; CIBERNED, Network Center for Biomedical Research in Neurodegenerative Diseases, National Institute of Health Carlos III, Madrid, Spain
| | - Silvia Gil
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain
| | - Miguel Ángel Santos-Santos
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain
| | - Ana Espinosa
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain; CIBERNED, Network Center for Biomedical Research in Neurodegenerative Diseases, National Institute of Health Carlos III, Madrid, Spain
| | - Gemma Ortega
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain; CIBERNED, Network Center for Biomedical Research in Neurodegenerative Diseases, National Institute of Health Carlos III, Madrid, Spain
| | - Ángela Sanabria
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain; CIBERNED, Network Center for Biomedical Research in Neurodegenerative Diseases, National Institute of Health Carlos III, Madrid, Spain
| | - Alba Pérez-Cordón
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain
| | - Pilar Cañabate
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain; CIBERNED, Network Center for Biomedical Research in Neurodegenerative Diseases, National Institute of Health Carlos III, Madrid, Spain
| | - Mariola Moreno
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain
| | - Silvia Preckler
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain
| | - Susana Ruiz
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain; CIBERNED, Network Center for Biomedical Research in Neurodegenerative Diseases, National Institute of Health Carlos III, Madrid, Spain
| | - Nuria Aguilera
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain
| | - Juan Antonio Pineda
- Unidad Clínica de Enfermedades Infecciosas y Microbiología. Hospital Universitario de Valme, Sevilla, Spain
| | - Juan Macías
- Unidad Clínica de Enfermedades Infecciosas y Microbiología. Hospital Universitario de Valme, Sevilla, Spain
| | - Emilio Alarcón-Martín
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain; Department of Surgery, Biochemistry and Molecular Biology, School of Medicine, University of Málaga, Málaga, Spain
| | - Oscar Sotolongo-Grau
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain
| | | | | | | | - Marta Marquié
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain
| | - Gemma Monté-Rubio
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain
| | - Sergi Valero
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain; CIBERNED, Network Center for Biomedical Research in Neurodegenerative Diseases, National Institute of Health Carlos III, Madrid, Spain
| | - Alba Benaque
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain
| | - Jordi Clarimón
- CIBERNED, Network Center for Biomedical Research in Neurodegenerative Diseases, National Institute of Health Carlos III, Madrid, Spain; Memory Unit, Neurology Department and Sant Pau Biomedical Research Institute, Hospital de la Santa Creu i Sant Pau, Universitat Autònoma de Barcelona, Barcelona, Spain
| | - Maria Jesus Bullido
- CIBERNED, Network Center for Biomedical Research in Neurodegenerative Diseases, National Institute of Health Carlos III, Madrid, Spain; Centro de Biologia Molecular Severo Ochoa (C.S.I.C.-U.A.M.), Universidad Autonoma de Madrid, Madrid, Spain; Instituto de Investigacion Sanitaria "Hospital la Paz" (IdIPaz), Madrid, Spain
| | | | - Pau Pástor
- Fundació per la Recerca Biomèdica i Social Mútua Terrassa, and Memory Disorders Unit, Department of Neurology, Hospital Universitari Mutua de Terrassa, University of Barcelona School of Medicine, Terrassa, Spain
| | - Pascual Sánchez-Juan
- CIBERNED, Network Center for Biomedical Research in Neurodegenerative Diseases, National Institute of Health Carlos III, Madrid, Spain; Neurology Service 'Marqués de Valdecilla' University Hospital (University of Cantabria and IDIVAL), Santander, Spain
| | - Victoria Álvarez
- Laboratorio de Genética Hospital Universitario Central de Asturias, Oviedo, Spain; Instituto de Investigación Biosanitaria del Principado de Asturias (ISPA), Oviedo, Spain
| | - Gerard Piñol-Ripoll
- CIBERNED, Network Center for Biomedical Research in Neurodegenerative Diseases, National Institute of Health Carlos III, Madrid, Spain; Unitat Trastorns Cognitius, Hospital Universitari Santa Maria de Lleida, Institut de Recerca Biomédica de Lleida (IRBLLeida), Lleida, Spain
| | | | - José Luis Royo
- Department of Surgery, Biochemistry and Molecular Biology, School of Medicine, University of Málaga, Málaga, Spain
| | - Emilio Franco
- Unidad de Demencias, Servicio de Neurología y Neurofisiología. Instituto de Biomedicina de Sevilla (IBiS), Hospital Universitario Virgen del Rocío/CSIC/Universidad de Sevilla, Seville, Spain
| | - Pablo Mir
- CIBERNED, Network Center for Biomedical Research in Neurodegenerative Diseases, National Institute of Health Carlos III, Madrid, Spain; Unidad de Trastornos del Movimiento, Servicio de Neurología y Neurofisiología. Instituto de Biomedicina de Sevilla (IBiS), Hospital Universitario Virgen del Rocío/CSIC/Universidad de Sevilla, Seville, Spain
| | - Miguel Calero
- CIBERNED, Network Center for Biomedical Research in Neurodegenerative Diseases, National Institute of Health Carlos III, Madrid, Spain; CIEN Foundation, Queen Sofia Foundation Alzheimer Center, Madrid, Spain; Instituto de Salud Carlos III (ISCIII), Madrid, Spain
| | - Miguel Medina
- CIBERNED, Network Center for Biomedical Research in Neurodegenerative Diseases, National Institute of Health Carlos III, Madrid, Spain; CIEN Foundation, Queen Sofia Foundation Alzheimer Center, Madrid, Spain
| | - Alberto Rábano
- CIBERNED, Network Center for Biomedical Research in Neurodegenerative Diseases, National Institute of Health Carlos III, Madrid, Spain; CIEN Foundation, Queen Sofia Foundation Alzheimer Center, Madrid, Spain; BT-CIEN, Madrid, Spain
| | - Jesús Ávila
- CIBERNED, Network Center for Biomedical Research in Neurodegenerative Diseases, National Institute of Health Carlos III, Madrid, Spain; Department of Molecular Neuropathology, Centro de Biología Molecular "Severo Ochoa" (CBMSO), Consejo Superior de Investigaciones Científicas (CSIC)/Universidad Autónoma de Madrid (UAM), Madrid, Spain
| | - Carmen Antúnez
- Unidad de Demencias, Hospital Clínico Universitario Virgen de la Arrixaca, Murcia, Spain
| | - Luis Miguel Real
- Unidad Clínica de Enfermedades Infecciosas y Microbiología. Hospital Universitario de Valme, Sevilla, Spain; Department of Surgery, Biochemistry and Molecular Biology, School of Medicine, University of Málaga, Málaga, Spain
| | - Adelina Orellana
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain
| | - Ángel Carracedo
- Grupo de Medicina Xenómica, Centro Nacional de Genotipado (CEGEN-PRB3-ISCIII). Universidade de Santiago de Compostela, Santiago de Compostela, Spain; Fundación Pública Galega de Medicina Xenómica- CIBERER-IDIS, Santiago de Compostela, Spain
| | | | - Lluís Tárraga
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain; CIBERNED, Network Center for Biomedical Research in Neurodegenerative Diseases, National Institute of Health Carlos III, Madrid, Spain
| | - Mercè Boada
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain; CIBERNED, Network Center for Biomedical Research in Neurodegenerative Diseases, National Institute of Health Carlos III, Madrid, Spain
| | - Agustín Ruiz
- Research Center and Memory clinic Fundació ACE, Institut Català de Neurociències Aplicades, Universitat Internacional de Catalunya, Barcelona, Spain; CIBERNED, Network Center for Biomedical Research in Neurodegenerative Diseases, National Institute of Health Carlos III, Madrid, Spain.
| |
Collapse
|
|
36
|
Tong SJ, Wall AA, Hung Y, Luo L, Stow JL. Guanine nucleotide exchange factors activate Rab8a for Toll-like receptor signalling. Small GTPases 2019; 12:27-43. [PMID: 30843452 DOI: 10.1080/21541248.2019.1587278] [Citation(s) in RCA: 13] [Impact Index Per Article: 2.6] [Reference Citation Analysis] [Abstract] [Key Words] [Track Full Text] [Journal Information] [Subscribe] [Scholar Register] [Indexed: 02/03/2023] Open
Abstract
Macrophages are important immune sentinels that detect and clear pathogens and initiate inflammatory responses through the activation of surface receptors, including Toll-like receptors (TLRs). Activated TLRs employ complex cellular trafficking and signalling pathways to initiate transcription for inflammatory cytokine programs. We have previously shown that Rab8a is activated by multiple TLRs and regulates downstream Akt/mTOR signalling by recruiting the effector PI3Kγ, but the guanine nucleotide exchange factors (GEF) canonically required for Rab8a activation in TLR pathways is not known. Using GST affinity pull-downs and mass spectrometry analysis, we identified a Rab8 specific GEF, GRAB, as a Rab8a binding partner in LPS-activated macrophages. Co-immunoprecipitation and fluorescence microscopy showed that both GRAB and a structurally similar GEF, Rabin8, undergo LPS-inducible binding to Rab8a and are localised on cell surface ruffles and macropinosomes where they coincide with sites of Rab8a mediated signalling. Rab nucleotide activation assays with CRISPR-Cas9 mediated knock-out (KO) cell lines of GRAB, Rabin8 and double KOs showed that both GEFs contribute to TLR4 induced Rab8a GTP loading, but not membrane recruitment. In addition, measurement of signalling profiles and live cell imaging with the double KOs revealed that either GEF is individually sufficient to mediate PI3Kγ-dependent Akt/mTOR signalling at macropinosomes during TLR4-driven inflammation, suggesting a redundant relationship between these proteins. Thus, both GRAB and Rabin8 are revealed as key positive regulators of Rab8a nucleotide exchange for TLR signalling and inflammatory programs. These GEFs may be useful as potential targets for manipulating inflammation. Abbreviations: TLR: Toll-like Receptor; OCRL: oculocerebrorenal syndrome of Lowe protein; PI3Kγ: phosphoinositol-3-kinase gamma; LPS: lipopolysaccharide; GEF: guanine nucleotide exchange factor; GST: glutathione S-transferases; BMMs: bone marrow derived macrophages; PH: pleckstrin homology; GAP: GTPase activating protein; ABCA1: ATP binding cassette subfamily A member 1; GDI: GDP dissociation inhibitor; LRP1: low density lipoprotein receptor-related protein 1.
Collapse
Affiliation(s)
- Samuel J Tong
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research (CIDR), The University of Queensland , Brisbane, QLD, Australia
| | - Adam A Wall
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research (CIDR), The University of Queensland , Brisbane, QLD, Australia
| | - Yu Hung
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research (CIDR), The University of Queensland , Brisbane, QLD, Australia
| | - Lin Luo
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research (CIDR), The University of Queensland , Brisbane, QLD, Australia
| | - Jennifer L Stow
- Institute for Molecular Bioscience (IMB) and IMB Centre for Inflammation and Disease Research (CIDR), The University of Queensland , Brisbane, QLD, Australia
| |
Collapse
|
|
37
|
Smith M, García-Martínez E, Pitter MR, Fucikova J, Spisek R, Zitvogel L, Kroemer G, Galluzzi L. Trial Watch: Toll-like receptor agonists in cancer immunotherapy. Oncoimmunology 2018; 7:e1526250. [PMID: 30524908 DOI: 10.1080/2162402x.2018.1526250] [Citation(s) in RCA: 162] [Impact Index Per Article: 27.0] [Reference Citation Analysis] [Abstract] [Key Words] [Track Full Text] [Journal Information] [Subscribe] [Scholar Register] [Received: 09/04/2018] [Indexed: 12/14/2022] Open
Abstract
Toll-like receptor (TLR) agonists demonstrate therapeutic promise as immunological adjuvants for anticancer immunotherapy. To date, three TLR agonists have been approved by US regulatory agencies for use in cancer patients. Additionally, the potential of hitherto experimental TLR ligands to mediate clinically useful immunostimulatory effects has been extensively investigated over the past few years. Here, we summarize recent preclinical and clinical advances in the development of TLR agonists for cancer therapy.
Collapse
Affiliation(s)
- Melody Smith
- Department of Medicine and Immunology Program, Memorial Sloan Kettering Cancer Center, New York, NY, USA
| | - Elena García-Martínez
- Hematology and Oncology Department, Hospital Universitario Morales Meseguer, Murcia, Spain
| | - Michael R Pitter
- Department of Medicine and Immunology Program, Memorial Sloan Kettering Cancer Center, New York, NY, USA
| | - Jitka Fucikova
- Sotio a.c., Prague, Czech Republic.,Department of Immunology, 2nd Faculty of Medicine, University Hospital Motol, Charles University, Prague, Czech Republic
| | - Radek Spisek
- Sotio a.c., Prague, Czech Republic.,Department of Immunology, 2nd Faculty of Medicine, University Hospital Motol, Charles University, Prague, Czech Republic
| | - Laurence Zitvogel
- INSERM, U1015, Villejuif, France.,Gustave Roussy Comprehensive Cancer Institute, Villejuif, France.,Center of Clinical Investigations in Biotherapies of Cancer (CICBT) 1428, Villejuif, France.,Université Paris Sud/Paris XI, Le Kremlin-Bicêtre, France
| | - Guido Kroemer
- Université Paris Descartes/ Paris V, Paris, France.,Université Pierre et Marie Curie/Paris VI, Paris, France.,INSERM, U1138, Paris, France.,Equipe 11 labellisée Ligue contre le Cancer, Centre de Recherche des Cordeliers, Paris, France.,Metabolomics and Cell Biology Platforms, Gustave Roussy Comprehensive Cancer Institute, Villejuif, France.,Karolinska Institute, Department of Women's and Children's Health, Karolinska University Hospital, Stockholm, Sweden.,Pôle de Biologie, Hopitâl Européen George Pompidou, AP-HP; Paris, France
| | - Lorenzo Galluzzi
- Université Paris Descartes/ Paris V, Paris, France.,Department of Radiation Oncology, Weill Cornell Medical College, New York, NY, USA.,Sandra and Edward Meyer Cancer Center, New York, NY, USA
| |
Collapse
|
|
38
|
Katikaneni DS, Jin L. B cell MHC class II signaling: A story of life and death. Hum Immunol 2018; 80:37-43. [PMID: 29715484 DOI: 10.1016/j.humimm.2018.04.013] [Citation(s) in RCA: 24] [Impact Index Per Article: 4.0] [Reference Citation Analysis] [Abstract] [Key Words] [Track Full Text] [Journal Information] [Subscribe] [Scholar Register] [Received: 01/31/2018] [Revised: 04/08/2018] [Accepted: 04/25/2018] [Indexed: 01/17/2023]
Abstract
MHC class II regulates B cell activation, proliferation, and differentiation during cognate B cell-T cell interaction. This is, in part, due to the MHC class II signaling in B cells. Activation of MHC Class II in human B cells or "primed" murine B cells leads to tyrosine phosphorylation, calcium mobilization, AKT, ERK, JNK activation. In addition, crosslinking MHC class II with monoclonal Abs kill malignant human B cells. Several humanized anti-HLA-DR/MHC class II monoclonal Abs entered clinical trials for lymphoma/leukemia and MHC class II-expressing melanomas. Mechanistically, MHC class II is associated with a wealth of transmembrane proteins including the B cell-specific signaling proteins CD79a/b, CD19 and a group of four-transmembrane proteins including tetraspanins and the apoptotic protein MPYS/STING. Furthermore, MHC class II signals are compartmentalized in the tetraspanin-enriched microdomains. In this review, we discuss our current understanding of MHC class II signaling in B cells focusing on its physiological significance and the therapeutic potential.
Collapse
Affiliation(s)
- Divya Sai Katikaneni
- Division of Pulmonary, Critical Care and Sleep Medicine, Department of Medicine, University of Florida, Gainesville, FL 32610, United States
| | - Lei Jin
- Division of Pulmonary, Critical Care and Sleep Medicine, Department of Medicine, University of Florida, Gainesville, FL 32610, United States.
| |
Collapse
|
|
39
|
Curson JEB, Luo L, Sweet MJ, Stow JL. pTRAPs: Transmembrane adaptors in innate immune signaling. J Leukoc Biol 2018; 103:1011-1019. [PMID: 29601097 DOI: 10.1002/jlb.2ri1117-474r] [Citation(s) in RCA: 9] [Impact Index Per Article: 1.5] [Reference Citation Analysis] [Abstract] [Key Words] [Track Full Text] [Journal Information] [Subscribe] [Scholar Register] [Received: 11/30/2017] [Revised: 02/08/2018] [Accepted: 02/10/2018] [Indexed: 01/30/2023] Open
Abstract
Transmembrane adaptor proteins (TRAPs) are protein scaffolds and signaling regulators with established roles in signal-induced activation of lymphocytes. A subset of the TRAP family, the palmitoylated TRAPs (pTRAPs), are increasingly emerging with additional roles in innate immune cells. Targeted to lipid rafts, tetraspannin-enriched microdomains, and protein microclusters in membranes, pTRAP scaffolds exert spatiotemporal regulation by recruiting signaling kinases, particularly Src and Syk family members, as well as Csk, and other effectors. In this way, pTRAPs modulate signaling and influence resulting cell responses, including the selective output of inflammatory cytokines and other mediators. Here, we review studies revealing that different pTRAPs work together, often with overlapping or redundant roles, for positive and negative regulation of key innate immune pathways, including Fc receptor and pattern recognition receptor signaling. Recent findings show that pTRAPs can bind directly to innate immune receptors, in addition to other transmembrane binding partners. Thus, pTRAPs are important, multifunctional scaffolds in pathways that are fundamental to diverse innate immune responses.
Collapse
Affiliation(s)
- James E B Curson
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research and Australian Infectious Diseases Research Centre, The University of Queensland, Brisbane, Queensland, Australia
| | - Lin Luo
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research and Australian Infectious Diseases Research Centre, The University of Queensland, Brisbane, Queensland, Australia
| | - Matthew J Sweet
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research and Australian Infectious Diseases Research Centre, The University of Queensland, Brisbane, Queensland, Australia
| | - Jennifer L Stow
- Institute for Molecular Bioscience (IMB), IMB Centre for Inflammation and Disease Research and Australian Infectious Diseases Research Centre, The University of Queensland, Brisbane, Queensland, Australia
| |
Collapse
|
|
40
|
Clark EL, Bush SJ, McCulloch MEB, Farquhar IL, Young R, Lefevre L, Pridans C, Tsang HG, Wu C, Afrasiabi C, Watson M, Whitelaw CB, Freeman TC, Summers KM, Archibald AL, Hume DA. A high resolution atlas of gene expression in the domestic sheep (Ovis aries). PLoS Genet 2017; 13:e1006997. [PMID: 28915238 PMCID: PMC5626511 DOI: 10.1371/journal.pgen.1006997] [Citation(s) in RCA: 92] [Impact Index Per Article: 13.1] [Reference Citation Analysis] [Abstract] [MESH Headings] [Grants] [Track Full Text] [Download PDF] [Figures] [Journal Information] [Subscribe] [Scholar Register] [Received: 06/08/2017] [Revised: 10/03/2017] [Accepted: 08/24/2017] [Indexed: 02/08/2023] Open
Abstract
Sheep are a key source of meat, milk and fibre for the global livestock sector, and an important biomedical model. Global analysis of gene expression across multiple tissues has aided genome annotation and supported functional annotation of mammalian genes. We present a large-scale RNA-Seq dataset representing all the major organ systems from adult sheep and from several juvenile, neonatal and prenatal developmental time points. The Ovis aries reference genome (Oar v3.1) includes 27,504 genes (20,921 protein coding), of which 25,350 (19,921 protein coding) had detectable expression in at least one tissue in the sheep gene expression atlas dataset. Network-based cluster analysis of this dataset grouped genes according to their expression pattern. The principle of 'guilt by association' was used to infer the function of uncharacterised genes from their co-expression with genes of known function. We describe the overall transcriptional signatures present in the sheep gene expression atlas and assign those signatures, where possible, to specific cell populations or pathways. The findings are related to innate immunity by focusing on clusters with an immune signature, and to the advantages of cross-breeding by examining the patterns of genes exhibiting the greatest expression differences between purebred and crossbred animals. This high-resolution gene expression atlas for sheep is, to our knowledge, the largest transcriptomic dataset from any livestock species to date. It provides a resource to improve the annotation of the current reference genome for sheep, presenting a model transcriptome for ruminants and insight into gene, cell and tissue function at multiple developmental stages.
Collapse
Affiliation(s)
- Emily L. Clark
- The Roslin Institute and Royal (Dick) School of Veterinary Studies, University of Edinburgh, Edinburgh, Scotland, United Kingdom
| | - Stephen J. Bush
- The Roslin Institute and Royal (Dick) School of Veterinary Studies, University of Edinburgh, Edinburgh, Scotland, United Kingdom
| | - Mary E. B. McCulloch
- The Roslin Institute and Royal (Dick) School of Veterinary Studies, University of Edinburgh, Edinburgh, Scotland, United Kingdom
| | - Iseabail L. Farquhar
- The Roslin Institute and Royal (Dick) School of Veterinary Studies, University of Edinburgh, Edinburgh, Scotland, United Kingdom
| | - Rachel Young
- The Roslin Institute and Royal (Dick) School of Veterinary Studies, University of Edinburgh, Edinburgh, Scotland, United Kingdom
| | - Lucas Lefevre
- The Roslin Institute and Royal (Dick) School of Veterinary Studies, University of Edinburgh, Edinburgh, Scotland, United Kingdom
| | - Clare Pridans
- The Roslin Institute and Royal (Dick) School of Veterinary Studies, University of Edinburgh, Edinburgh, Scotland, United Kingdom
| | - Hiu G. Tsang
- The Roslin Institute and Royal (Dick) School of Veterinary Studies, University of Edinburgh, Edinburgh, Scotland, United Kingdom
| | - Chunlei Wu
- Department of Integrative and Computational Biology, The Scripps Research Institute, La Jolla, CA, United States of America
| | - Cyrus Afrasiabi
- Department of Integrative and Computational Biology, The Scripps Research Institute, La Jolla, CA, United States of America
| | - Mick Watson
- The Roslin Institute and Royal (Dick) School of Veterinary Studies, University of Edinburgh, Edinburgh, Scotland, United Kingdom
| | - C. Bruce Whitelaw
- The Roslin Institute and Royal (Dick) School of Veterinary Studies, University of Edinburgh, Edinburgh, Scotland, United Kingdom
| | - Tom C. Freeman
- The Roslin Institute and Royal (Dick) School of Veterinary Studies, University of Edinburgh, Edinburgh, Scotland, United Kingdom
| | - Kim M. Summers
- The Roslin Institute and Royal (Dick) School of Veterinary Studies, University of Edinburgh, Edinburgh, Scotland, United Kingdom
- Mater Research Institute and University of Queensland, Translational Research Institute, Woolloongabba, Queensland, Australia
| | - Alan L. Archibald
- The Roslin Institute and Royal (Dick) School of Veterinary Studies, University of Edinburgh, Edinburgh, Scotland, United Kingdom
| | - David A. Hume
- The Roslin Institute and Royal (Dick) School of Veterinary Studies, University of Edinburgh, Edinburgh, Scotland, United Kingdom
- Mater Research Institute and University of Queensland, Translational Research Institute, Woolloongabba, Queensland, Australia
| |
Collapse
|
|
41
|
Development of SH2 probes and pull‐down assays to detect pathogen‐induced, site‐specific tyrosine phosphorylation of the TLR adaptor SCIMP. Immunol Cell Biol 2017; 95:564-570. [DOI: 10.1038/icb.2017.10] [Citation(s) in RCA: 6] [Impact Index Per Article: 0.9] [Reference Citation Analysis] [Track Full Text] [Journal Information] [Subscribe] [Scholar Register] [Received: 01/09/2017] [Revised: 02/08/2017] [Accepted: 02/10/2017] [Indexed: 01/12/2023]
|